Lab 6: Multicollinearity, Dominance Analysis, and Power
PSYC 7804 - Regression with Lab
Spring 2025
Today’s Packages and Data 🤗
The dominanceanalysis package (Navarrete & Soares, 2024) contains functions to run dominance analysis in R.
The pwr package (Champely et al., 2020) includes functions to conduct power analysis for some statistical analyses and experimental designs. Click here for examples on all the types of power calculations that the package allows for.
Background About the data
I am using some of the data from Setti & Kahn (2024), where (yes I am citing myself, but it makes sense for this lab, I promise 😶🫣) we looked at how facets of openness to experience (both from the big five and the HEXACO) predict music preference.
For the data here I selected only 3 personality facets and 1 dimension of music preference.
- Intense: Preference for the intense music dimension, which includes genres such as rock, punk, metal.
- Advnt: Adventurousness facet of big 5 openness to experience.
- Intel: Intellect facet of big 5 openness to experience.
- Uncon: Unconventionality facet of HEXACO openness to experience.
Higher values mean higher preference/levels of personality trait
Dimensions of Music Preference 🎶
Rentfrow & Gosling (2003) was a seminal study in the field of music preference, where it was found that preference for music genres cluster together (e.g., if you like punk music, you tend to like metal music). Later Rentfrow et al. (2011) proposed a 5-factor structure of music preference: sophisticated (e.g., classical, blues, jazz), unpretentious (e.g., country and folk), intense (e.g, rock, punk, metal), mellow (e.g., pop, electronic), and contemporary (e.g., rap, RnB). This 5-factor structure seems to work fairly well, although I would suspect that 1 or 2 more factors would emerge with a more comprehensive selection of music genres.
This data fits the lab topic well because in the paper we had to figure out a way of dealing with high multicollinearity, and dominance analysis ended up helping a lot!
Regression coefficients Weirdness
Intense Advnt Intel
Intense 1.00 0.24 0.27
Advnt 0.24 1.00 0.96
Intel 0.27 0.96 1.00Advnt and Intense becomes negative after accounting for Intel 🙀
Advnt and Intense while accounting for Intel. We run a multiple regression:
Coefficients:
Estimate Std. Error t value Pr(>|t|)
(Intercept) -2.3076 0.4642 -4.972 1.02e-06 ***
Advnt -0.6444 0.4520 -1.426 0.1548
Intel 1.2766 0.4571 2.793 0.0055 **
---
Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
Residual standard deviation: 0.9686 on 366 degrees of freedom
Multiple R-squared: 0.07685
F-statistic: 15.23 on 2 and 366 DF, p-value: 4.416e-07
AIC BIC
1028.61 1044.26 Visualizing Multicollinearity
When strange things happen, visualizing your data (if possible) is always a good idea.
Advnt and Intel are extremely correlated (\(r = .96\)). You can see it from the 3D plot, which is essentially a 2D plot (Advnt and Intel are effectively a single variable, not two separate ones). This is a case of multicollinearity.
Back to Semi-Partial Correlations
If you remember, semi-partial correlations calculate the correlation between \(y\) and \(x\) after removing the explained variance of \(z\) from only one variable.
$estimate
Intense Advnt Intel
Intense 1.00000000 -0.0715963 0.1402480
Advnt -0.01976479 1.0000000 0.9340608
Intel 0.03841693 0.9268286 1.0000000We want to look at the first row. We can see that the relation between Intense and advnt is negative after the variance explained by Intel is taken out of Advnt.
As we saw a few slides back, by themselves both variables are positively correlated with Intense
Intense Advnt Intel
1.0000000 0.2391202 0.2678095 A Word of Advice
When the sign of correlations and semi-partial correlations are different, something strange is happening. Multicollinearity may be the reason, but other possibilities exist (see here).
Variance Inflation Factor
Probably the quickest way to check for multicollinearity is to calculate the variance inflation factor (VIF) for each predictor. For a predictor \(x\), the formula is:
\[VIF_x = \frac{1}{1 - R^2}\]
Importantly, the \(R^2\) in the VIF formula stands for the variance explained in the predictor \(x\) by all other predictors.
Another Perspective: Residuals
As shown in Lab 4, regression coefficients are calculated based on the residuals of the predictors after controlling for all other predictors. But, if Advnt and Intel are so correlated, what happens to their residual after controlling fo one another?
However, \(R^2\) is untouched by multicollinearity! So, you can still interpret \(R^2\) just fine with high multicollinearity.
Dominance Analysis
Dominance analysis (Azen & Budescu, 2003; Budescu, 1993) main application is to determine the relative importance of a set of predictor variables. DA provides a way of ranking predictors by importance.
Different dominance patterns (complete dominance, conditional dominance, general dominance) between variables can be established depending on the results of DA.

Example with our Data
* Fit index: r2
Average contribution of each variable:
Uncon Intel Advnt
0.064 0.034 0.023
Dominance Analysis matrix:
model level fit Advnt Intel Uncon
1 0 0 0.057 0.072 0.098
Advnt 1 0.057 0.02 0.054
Intel 1 0.072 0.005 0.045
Uncon 1 0.098 0.013 0.02
Average level 1 1 0.009 0.02 0.049
Advnt+Intel 2 0.077 0.044
Advnt+Uncon 2 0.111 0.011
Intel+Uncon 2 0.117 0.004
Average level 2 2 0.004 0.011 0.044
Advnt+Intel+Uncon 3 0.121 -
model: variables in the regression. -
level: number of variables in the regression. -
fit: \(R^2\) value for given regression. -
remaining columns: contribution in \(R^2\) when adding variable in column.
let’s go over the meaning of the columns in the output
Uncon contributes the most \(R^2\) on average, \(.064\) (i.e., general dominance).
Additional Dominance Patterns
So, on average Uncon is the most important predictor or preference for Intense music. This is just on average across all possible regression. We can build dominance matrices with j rows and i columns.
A dominance matrix will contain a 1 if the variable in the row \(j\) dominates the one in column \(i\). You will see a .5 if the dominance pattern between two variables cannot be established.
In both cases, Uncon dominates the other two variables. We now have a really strong case to claim that Uncon is the most important predictor of preference for Intense music among our 3 openness facets.
Uncertainty and Inference in DA
As it is the case with any statistical procedure, our results are based on a sample, and there is uncertainty about our results (e.g., are they due to sampling error?). We can use bootstrap to check how often the dominance pattern replicates across every bootstrap sample (1000 in this case):
Dominance Analysis
==================
Fit index: r2
dominance i k Dij mDij SE.Dij Pij Pji Pnoij Rep
complete Advnt Intel 0 0.032 0.155 0.018 0.954 0.028 0.954
complete Advnt Uncon 0 0.072 0.193 0.013 0.869 0.118 0.869
complete Intel Uncon 0 0.159 0.301 0.073 0.756 0.171 0.756
conditional Advnt Intel 0 0.032 0.155 0.018 0.954 0.028 0.954
conditional Advnt Uncon 0 0.072 0.193 0.013 0.869 0.118 0.869
conditional Intel Uncon 0 0.159 0.301 0.073 0.756 0.171 0.756
general Advnt Intel 0 0.022 0.147 0.022 0.978 0.000 0.978
general Advnt Uncon 0 0.052 0.222 0.052 0.948 0.000 0.948
general Intel Uncon 0 0.148 0.355 0.148 0.852 0.000 0.852Skipping over some stuff, the column that we care most about is the rep column.
This column shows the proportion of bootstrap samples where the dominance pattern in the row was replicated. In general, Uncon seems to fairly consistently dominate all other variables.
DA and Multicollinearity
What is Power⚡?
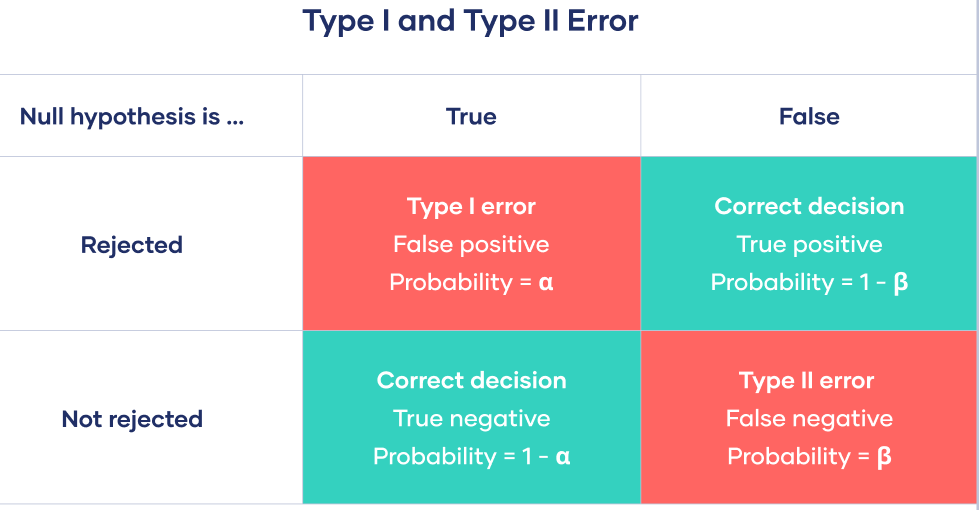
Power: Statistical power is defined as the probability of correctly rejecting the null hypothesis if \(H_0\) is false. Power is also known as 1 - Type II error rate. Type II error rate is denoted with \(\beta\), and is the probability of failing to reject a false \(H_0\).
Power for Correlation
Calculating power for correlation, \(r\), is fairly straightforward. In general calculating power involves these pieces of information:
The effect size. Could be \(r\), \(d\), \(f^2\), …
The significance level, \(\alpha\)
The power, \(1 - \beta\)
The sample size, \(N\)
pwr package works in the same way.
- If we expect to see \(r =.3\), what sample size do we need to have \(80\%\) power at \(\alpha = .05\)?
Power in Regression
In regression, We calculate power based on \(R^2\). More specifically, we use Cohen’s \(f^2\). there are two slight variations on \(f^2\):
When you have a single set of predictors and you want to calculate power for a regression,
\[ f^2 = \frac{R^2}{1 - R^2} \]
When you want to calculate power for \(\Delta R^2\) (see Lab 6), whether a set of predictors B provides a significant improvement over an original set of predictors A,
\[ f^2 = \frac{R^2_{AB} - R^2_{A}}{1 - R^2_{AB}} \] Where \(R^2_{AB} - R^2_{A} = \Delta R^2\), the improvement of adding the set of predictors B.
Larger \(R^2\) value imply larger \(f^2\) values. Some guidelines are that \(f^2\) of .02, .15, and .35 can be interpreted as small, medium, and large effect sizes respectively. Why do we use \(f^2\)? Check out the appendix if you are curios!
Power For \(R^2\)
u stands for \(\mathrm{df_1}\) and v stands for the \(\mathrm{df_2}\) of the \(F\)-distribution. We know from Lab 4 that \(\mathrm{df_2} = n - \mathrm{df_1} - 1\). Thus,
\[ n = \mathrm{df_2} + 1 + \mathrm{df_1} = 61.18 + 1 + 3 = 65.18 \]
Power For \(\Delta R^2\)
So, if we want to have \(80\%\) power for \(\Delta R^2 = .05\), we now need at least \(131\) participants. We know this because
\[ n = 125.5 + 4 + 1 = 130.5 \]
I already linked this in the first slide, but this page has very detailed explanations on how to use all the functions from the pwr package.
What about power for Slopes?
\[ \mathrm{SE}_{b_1} = \frac{S_Y}{S_{X_1}}\sqrt{\frac{1 - r_{Y\hat{Y}}}{n - p - 1}}\times\sqrt{\frac{1}{1 - r^2_{12}}} \]
Low Power is bad Because…
Low power is not bad because it will be hard to get \(p < .05\). Low power is bad because when you do get \(p < .05\) with low power, it is likely that:
- The regression coefficients will be largely overestimated! (Type M error).
- The sign of some regression coefficients will be in the wrong direction! (Type S error).
So, I think people care about power for the wrong reasons 🤷 Some related blog posts/manuscripts by Andrew Gelman:

References
Appendix: Why \(f^2\)? The non-central \(F\)-distribution
\(f^2\) and non-central \(F\)-distribution
We know that \(R^2\) is itself a measure of effect size; so why do we calculate \(f^2\) to compute power? Chapter 9 of good old Cohen (1988) goes in more detail, but \(f^2\) is more general and convenient than \(R^2\).
Power for \(F\)-tests is calculated by comparing two things:
- The null \(F\)-distribution if \(R^2 = 0\). This is a normal \(F\)-distribution with \(\mathrm{df_1} = k\) and \(\mathrm{df_2} = n - k-1\).
- The \(F\)-distribution if \(R^2 \neq 0\). This is an \(F\)-distribution with \(\mathrm{df_1} = k\), \(\mathrm{df_2} = n - k-1\), and a non-centrality parameter that we will call \(L\) (in Cohen (1988), this is referred to as \(\lambda\)).
The formula to calculate \(L\) is very straightforward once we have \(f^2\):
\[ L = f^2\times(\mathrm{df_1} + \mathrm{df_2} + 1) = f^2 \times n \]
Note that the sample size, \(n\), is included in this formula, and we can solve for it such that \(n = \frac{L}{f^2}\) (neat).
The Non-central F-distribution
Power by hand
oh, and because \(n = \mathrm{df_1} + \mathrm{df_2} + 1\), you can do some algebra on the \(L = f^2 \times n\) equation to find any of the other terms if you want.
Checking with the pwr.f2.test() function
As always, I want to check that the result I got is also what you would get with another function. Let’s use the pwr.f2.test():
Now I know with really high certainty that my understanding of the process is accurate.
PSYC 7804 - Lab 6: Multicollinearity, Dominance Analysis, and Power