Lab 8: Quadratic regression and non-linear alternatives
PSYC 7804 - Regression with Lab
Spring 2025
Today’s Packages and Data 🤗
The lubridate package (Spinu et al., 2024) provides many functions to help you work with date and times.
The splines2 package (Wang et al., 2024) can be used to calculate splines.
'data.frame': 59760 obs. of 10 variables:
$ time : chr "2016-01-01T00:00" "2016-01-01T01:00" ...
$ temperature_2m (°C) : num 7.6 7.5 7.1 6.6 6.3 ...
$ precipitation (mm) : num 0 0 0 0 0 ...
$ rain (mm) : num 0 0 0 0 0 ...
$ cloudcover (%) : num 69 20 32 35 34 ...
$ cloudcover_low (%) : num 53 4 3 5 4 ...
$ cloudcover_mid (%) : num 0 0 0 0 0 ...
$ cloudcover_high (%) : num 72 56 99 100 100 ...
$ windspeed_10m (km/h) : num 10 9.8 9.7 9.2 9.1 ...
$ winddirection_10m (°): num 296 287 285 281 279 ...The data includes weather recordings in NY for every hour of the day over a period of roughly 7 years (lots of data 😱). This is a bit too granular for what we need to do, so let’s do some data cleaning!
Working with Dates
This Data provides a good opportunity to show how you can work with dates in R! Although I will only show one function here, the lubridate package is your friend if you work a lot with dates and times in your research.
time variable into a date in days that R recognizes. we would use the ymd() function normally, but the correct format should be YYYY-MM-DD. Instead we also have the time here, which we do not need:
time variable into a date:
Summarizing and Filtering the Data
dplyr package, which is always loaded by tidyverse. Let’s get the average for each day:
tibble [2,490 × 2] (S3: tbl_df/tbl/data.frame)
$ time : Date[1:2490], format: "2016-01-01" "2016-01-02" ...
$ Temp_C: num [1:2490] 5.41 2.39 3.01 0.1 -6.78 ...The tidyverse Pipe Operator %>%
The %>% operator, AKA the pipe operator, passes the element on its left to the function to its right. It is helpful to do multiple operations without needing to redefine an object on every line.
tibble [364 × 3] (S3: tbl_df/tbl/data.frame)
$ time : Date[1:364], format: "2021-01-01" "2021-01-02" ...
$ Temp_C: num [1:364] 0.729 5.463 2.25 2.75 3.125 ...
$ Temp_F: num [1:364] 33.3 41.8 36 37 37.6 ...Plot The Data
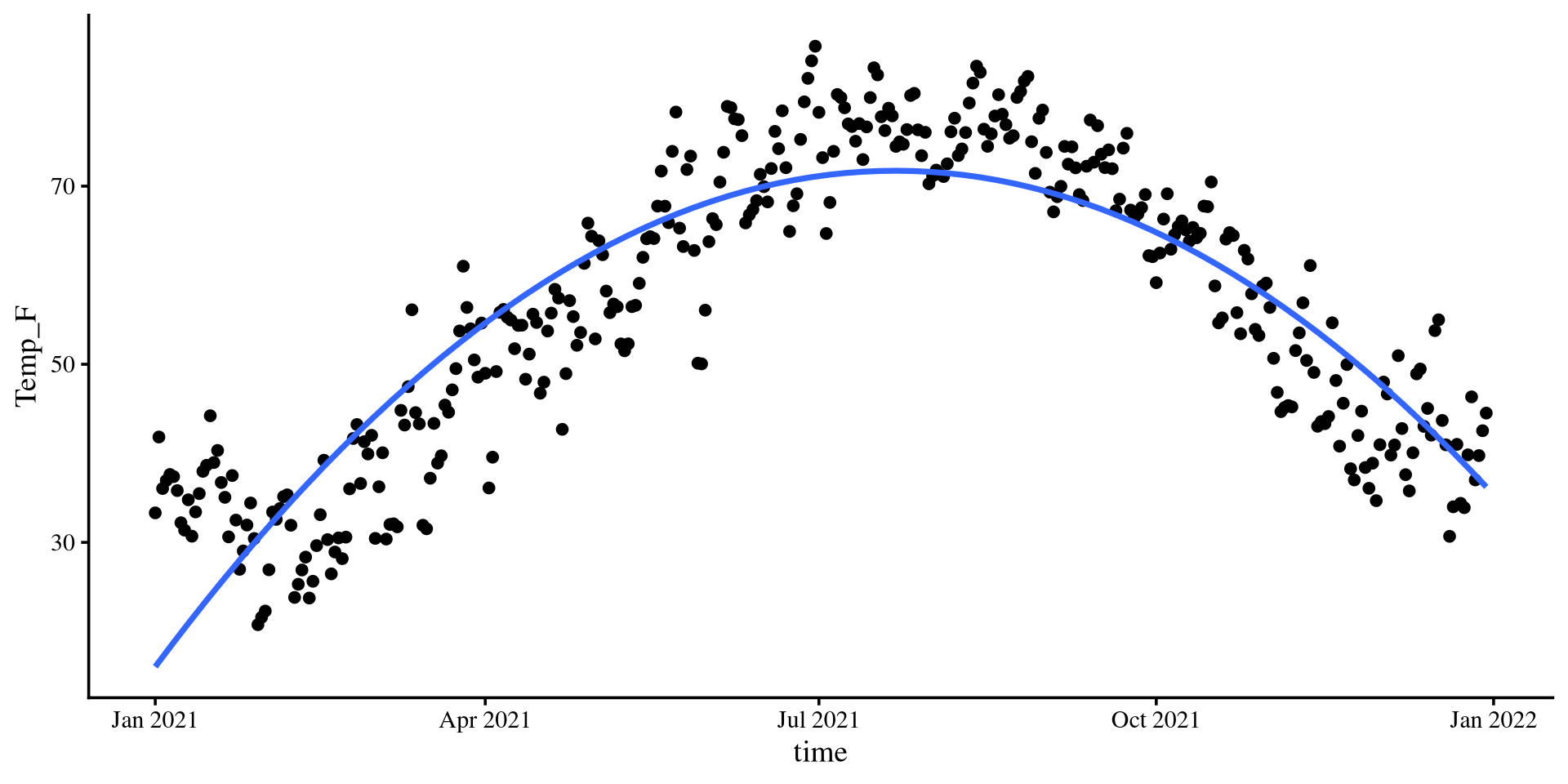
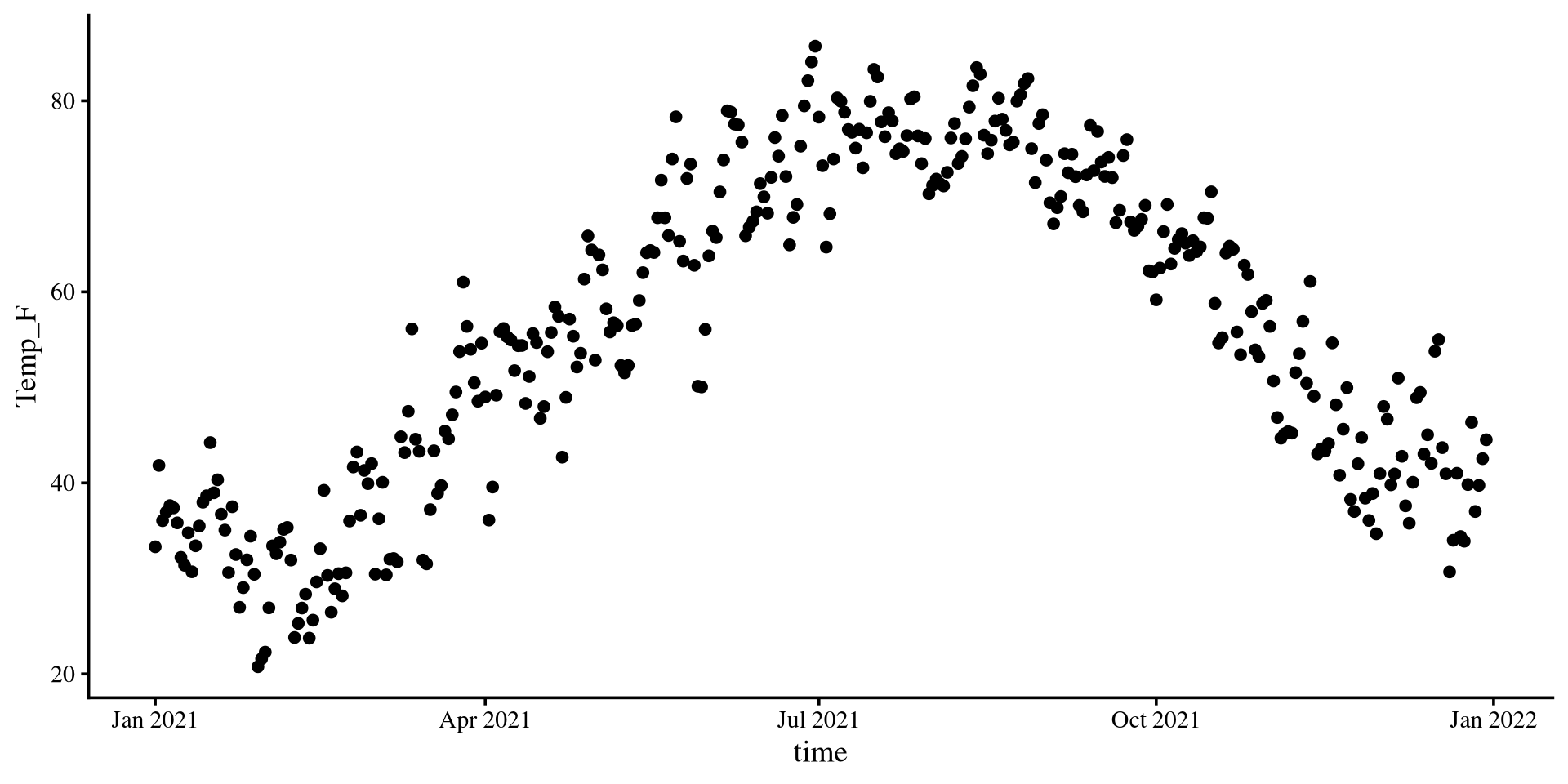
Let’s see what is the temperature trend in NY for the year 2021.
Plot The Data
Let’s see what is the temperature trend in NY for the year 2021.
Plot The Data
Let’s see what is the temperature trend in NY for the year 2021.
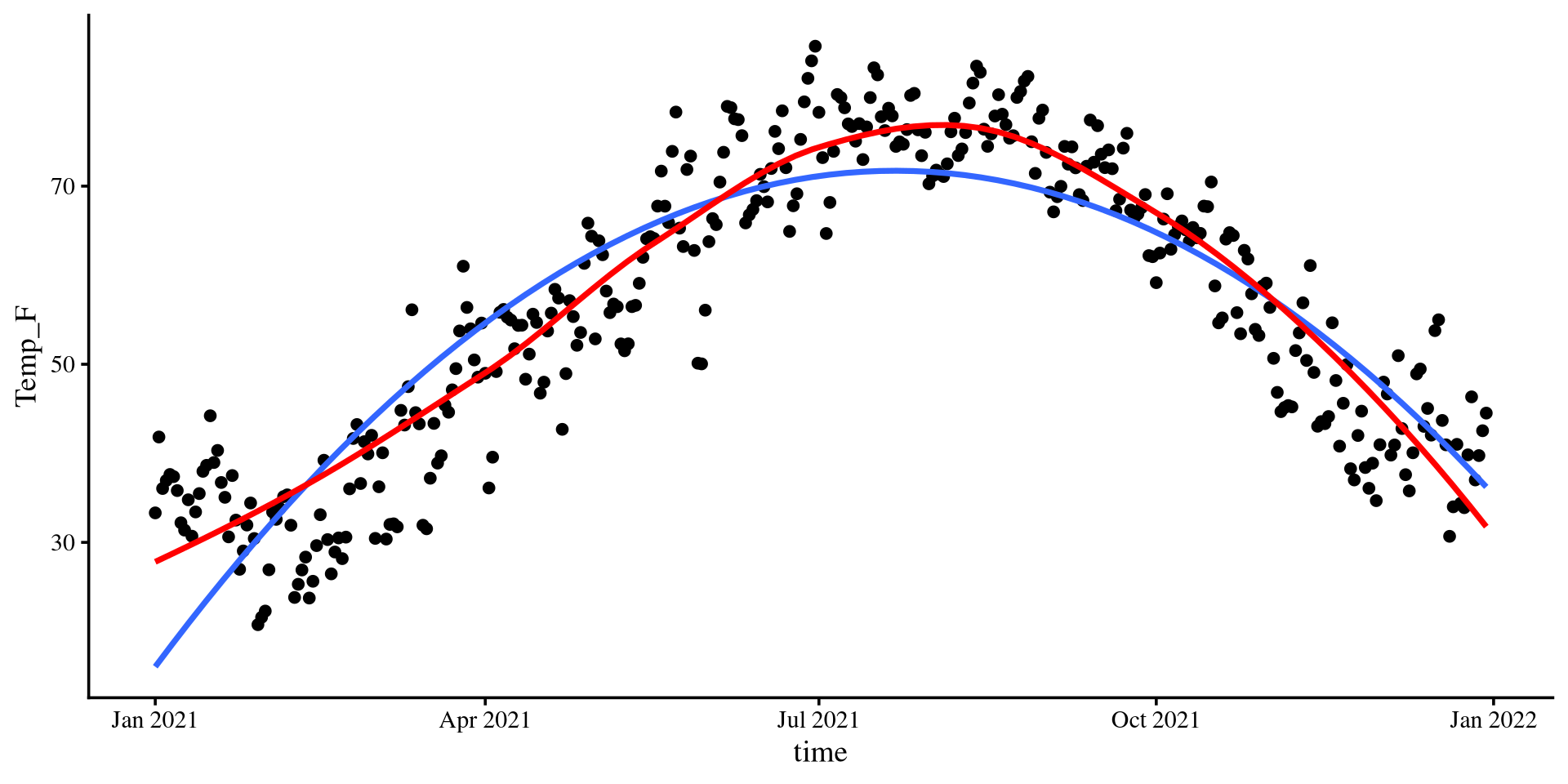
The loess line is pretty close to the quadratic one!
Let’s run a quadratic regression with time as the predictor…
Quadratic Regression: Centering Variables
time variable
We will not be able to interpret our regression coefficients (the linear term specifically) with such multicollinearity, as discussed in Lab 7
To center a variable we just subtract the mean
\[X_{\mathrm{centered}} = X - \bar{X}\]
And the mulitcollinearity problem is solved 🪄
Quadratic Regression
Call:
lm(formula = Temp_F ~ time_cent + time_cent2, data = Temp_avg_2021)
Residuals:
Min 1Q Median 3Q Max
-18.8248 -5.4369 0.3819 5.0293 25.2432
Coefficients:
Estimate Std. Error t value Pr(>|t|)
(Intercept) 7.116e+01 6.140e-01 115.90 <2e-16 ***
time_cent 5.562e-02 3.896e-03 14.28 <2e-16 ***
time_cent2 -1.367e-03 4.145e-05 -32.98 <2e-16 ***
---
Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
Residual standard error: 7.81 on 361 degrees of freedom
Multiple R-squared: 0.7815, Adjusted R-squared: 0.7803
F-statistic: 645.7 on 2 and 361 DF, p-value: < 2.2e-16\(\widehat{\mathrm{F_{temp}}} = b_0 + b_1 \times \mathrm{time} + b_2 \times \mathrm{time}^2\)
\(\widehat{\mathrm{F_{temp}}} = 71.16 + 0.06 \times \mathrm{time} - 0.001 \times \mathrm{time}^2\)
-
71.16 is the expected temperature in Fahrenheit when
timeis 0. Our variable is centered, so 0 represent the middle of the year, roughly July 2nd -
0.06 is the expected slope of the tangent line when
timeis 0 (confusing 😕) -
-0.001 is the expected change in the slope of the tangent line per unit-change in
time(even more confusing 😕😕)
We need some graphical help because some of this stuff honestly makes no sense in words 🤷
\(b_0\) in Quadratic Regression
Plot code
ggplot(Temp_avg_2021,
aes(x = time, y = Temp_F)) +
geom_point(alpha = .5, shape = 1) +
geom_point( aes(y = coef(quad_reg)[1],
x = mean(Temp_avg_2021$time)),
size = 3.5,
col = "red") +
geom_smooth(method = "lm",
formula = y ~ poly(x, 2),
se = FALSE) +
geom_segment(x = 0,
xend = mean(Temp_avg_2021$time),
y = coef(quad_reg)[1],
yend = coef(quad_reg)[1],
linetype = 2) +
geom_segment(x = mean(Temp_avg_2021$time),
xend = mean(Temp_avg_2021$time),
y = 0,
yend = coef(quad_reg)[1],
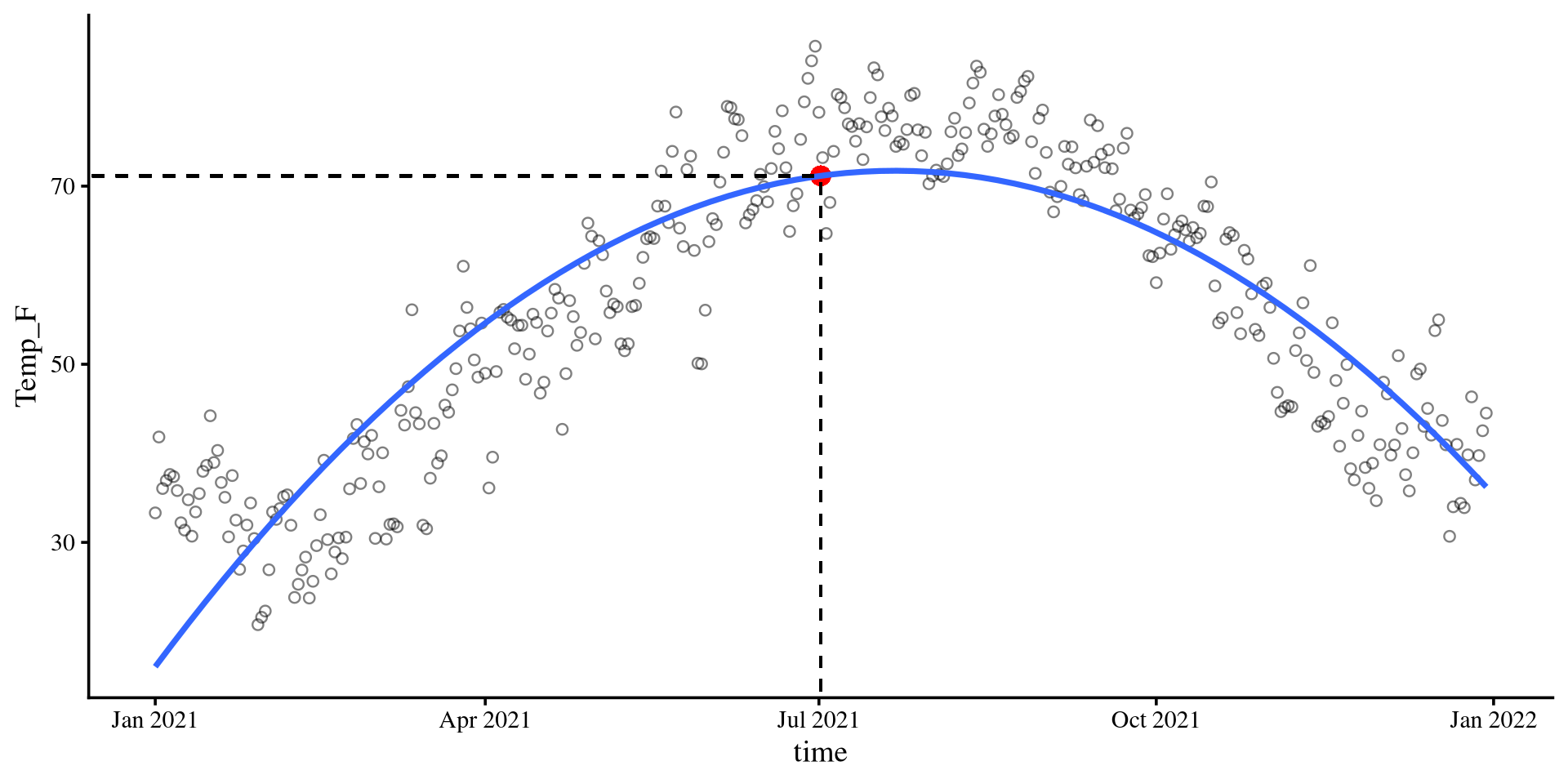
linetype = 2)\[b_0 = 71.16\]
Given that the time variable was centered, 0 corresponds to the middle of the year, July 2nd.
time = 0) given the quadratic regression equation is indeed 71.16 degrees Fahrenheit.
\(b_1\) in Quadratic Regression
Plot code
ggplot(Temp_avg_2021,
aes(x = time, y = Temp_F)) +
geom_point(alpha = .5, shape = 1) +
geom_smooth(method = "lm",
formula = y ~ poly(x, 2),
se = FALSE) +
geom_segment(x = mean(Temp_avg_2021$time) - 130,
xend = mean(Temp_avg_2021$time) + 130,
y = coef(quad_reg)[1] - coef(quad_reg)[2]*130,
yend = coef(quad_reg)[1] + coef(quad_reg)[2]*130,
linetype = 1,
linewidth = .7,
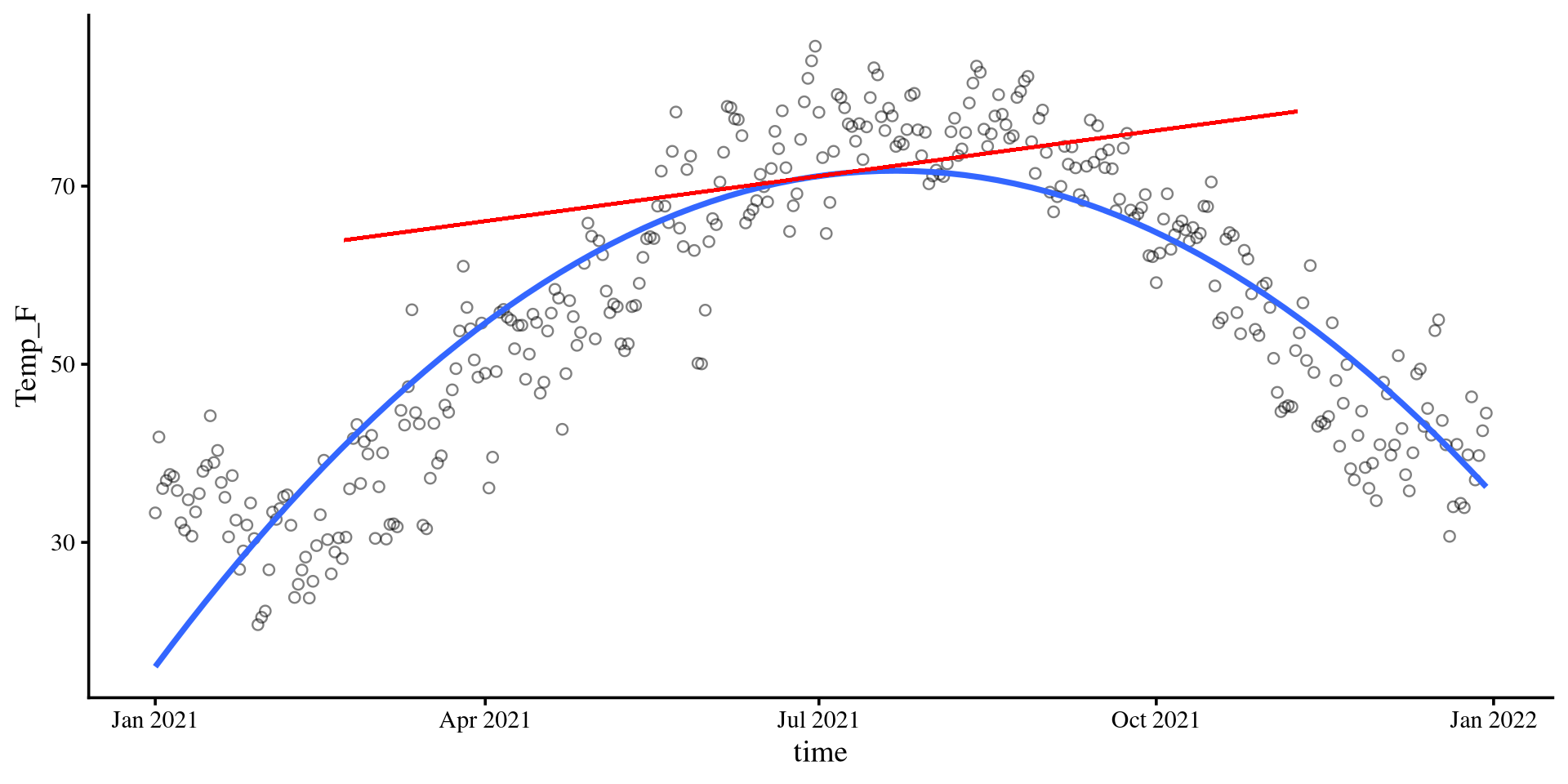
col = "red")\[b_1 = 0.06\]
The slope of the tangent line when time is at 0. The tangent line is the red line in the plot on the right.
Slope-ception 🤔
If you think about it, in linear regression, the slope, \(b_1\), is also the slope of the tangent line to the regression line.
\(b_2\) in Quadratic Regression
Plot code
x_val <- 100
# accurate calculations, there is rounding error in the equation on the slides
slope_new <- coef(quad_reg)[2] + 2*coef(quad_reg)[3]*x_val
ggplot(Temp_avg_2021,
aes(x = time, y = Temp_F)) +
geom_point(alpha = .5, shape = 1) +
geom_smooth(method = "lm",
formula = y ~ poly(x, 2),
se = FALSE) +
geom_segment(x = (mean(Temp_avg_2021$time) + x_val) - 130,
xend = (mean(Temp_avg_2021$time) + x_val) + 130,
y = (coef(quad_reg)[1] + coef(quad_reg)[2]*x_val + coef(quad_reg)[3]*x_val^2) - (slope_new*130),
yend = (coef(quad_reg)[1] + coef(quad_reg)[2]*x_val + coef(quad_reg)[3]*x_val^2) + (slope_new*130),
linetype = 1,
linewidth = .7,
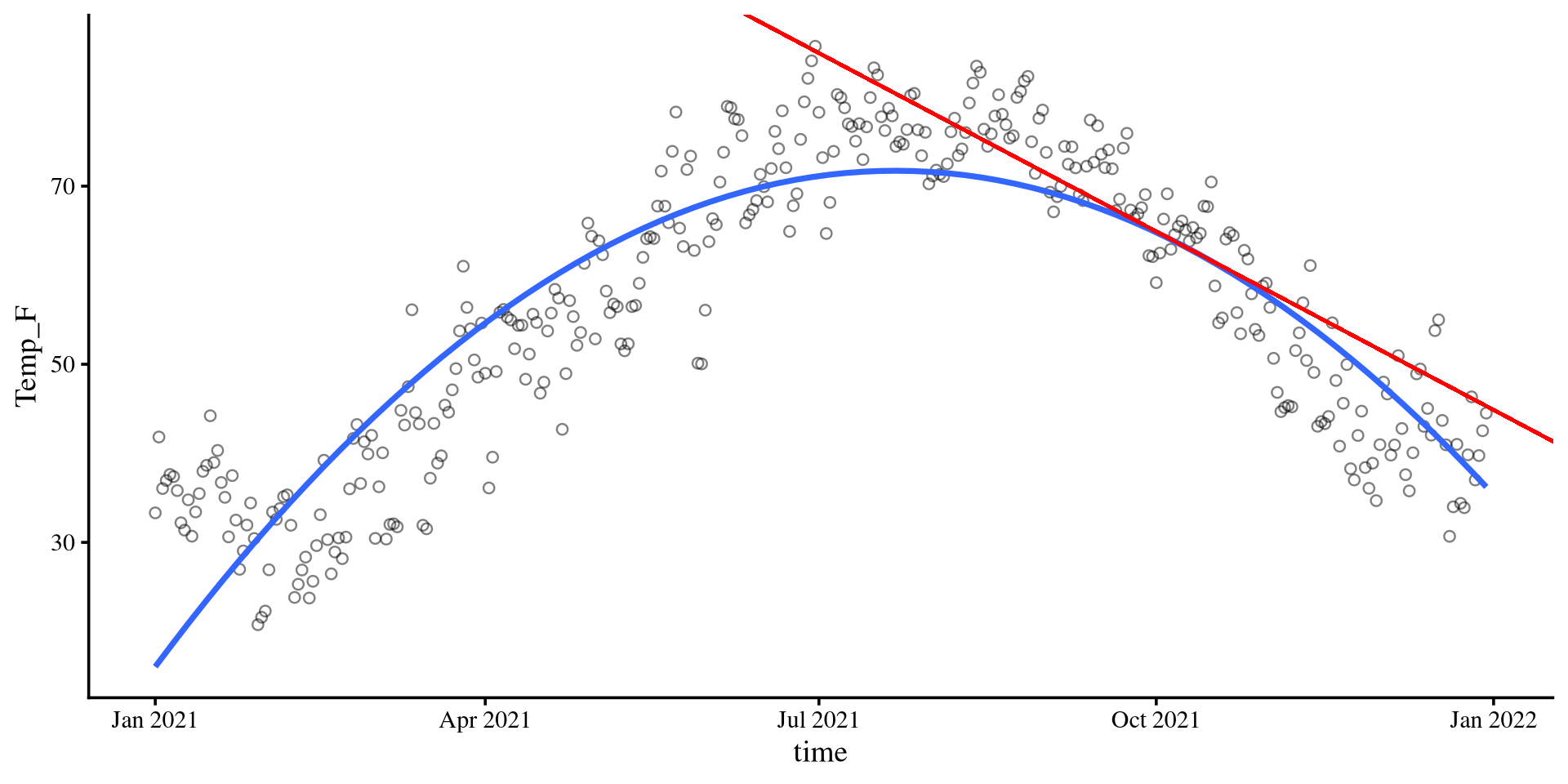
col = "red") \[b_2 = - 0.001\]
\(b_2\) lets us calculate the slope of the tangent line at any point of the curve. The value of \(b_2\), describes the change of the slope of the tangent line per 1-unit change in time. More formally,
time = 100, then the slope of the tangent line is given by \(b_{\mathrm{tang}} = .06 + 2\times -.001 \times100 = -.14\). Indeed, the tangent line goes downwards, as its slope suggests.
Why \(b_2\) is What You care about
So why do we care about slopes of tangent lines? Well, generally, we don’t. The thing that we care about is that the slope of tangent lines changes at all. This is what a non-zero \(b_2\) value tells us.
What would it mean for the slope of the tangent line to be constant across every \(X\) value? What happens if \(b_2 = 0\)?
We can check our quadratic regression equation, so \(\hat{Y} = b_0 + b_1X + 0X^2\) → \(\hat{Y} = b_0 + b_1X\)
Oh, we are back to the linear regression formula 🙃 If the slope of the tangent line is always constant, we have a line.
So, \(b_2\) informs us about 3 things:
1. Whether the line bends at all (i.e., is \(b_2 \neq 0\))
2. How much the line bends, which it depends on the magnitude of \(b_2\) (more extreme \(b_2\) values imply steeper curves)
3. The direction of the bend. If the sign of \(b_2\) is positive, the curve will be a bell, if the sign is negative, it will be a U, as in our case.
Climbing to the top ☝️ (or bottom 👇)
Plot code
# accurate calculations, there is rounding error in the equation on the slides
top <- -1*(coef(quad_reg)[2]/(2*coef(quad_reg)[3]))
Y_val <- (coef(quad_reg)[1] + coef(quad_reg)[2]*top + coef(quad_reg)[3]*top^2)
# this is the formula for the tangent slope. if you print this, you will see that it equals exactly 0. This means that the X value provided is the top of the curve, which is the only point where the tangent slope is exactly 0
slope_new <- coef(quad_reg)[2] + 2*coef(quad_reg)[3]*top
ggplot(Temp_avg_2021,
aes(x = time, y = Temp_F)) +
geom_point(alpha = .5, shape = 1) +
geom_point(aes(y = Y_val,
x = mean(Temp_avg_2021$time) + top),
size = 3.5,
col = "red") +
geom_smooth(method = "lm",
formula = y ~ poly(x, 2),
se = FALSE) +
geom_segment(x = 0,
xend = mean(Temp_avg_2021$time) + top,
y = Y_val,
yend = Y_val,
linetype = 2) +
geom_segment(x = mean(Temp_avg_2021$time) + top,
xend = mean(Temp_avg_2021$time) + top,
y = 0,
yend = Y_val,
linetype = 2) +
geom_segment(x = mean(Temp_avg_2021$time) -130,
xend = mean(Temp_avg_2021$time) + 130,
y = Y_val,
yend = Y_val,
linetype = 1,
linewidth = .7,
col = "red") 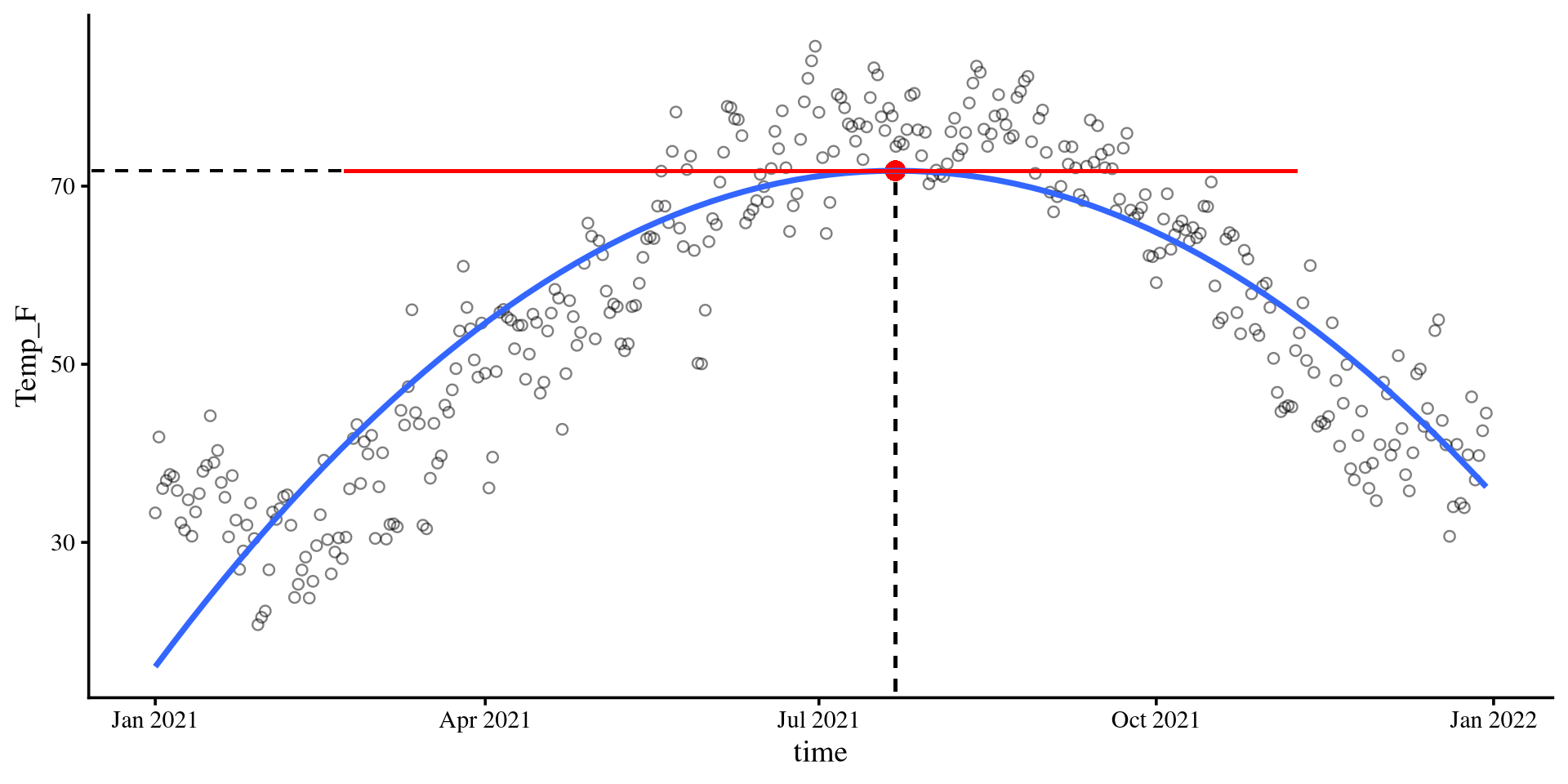
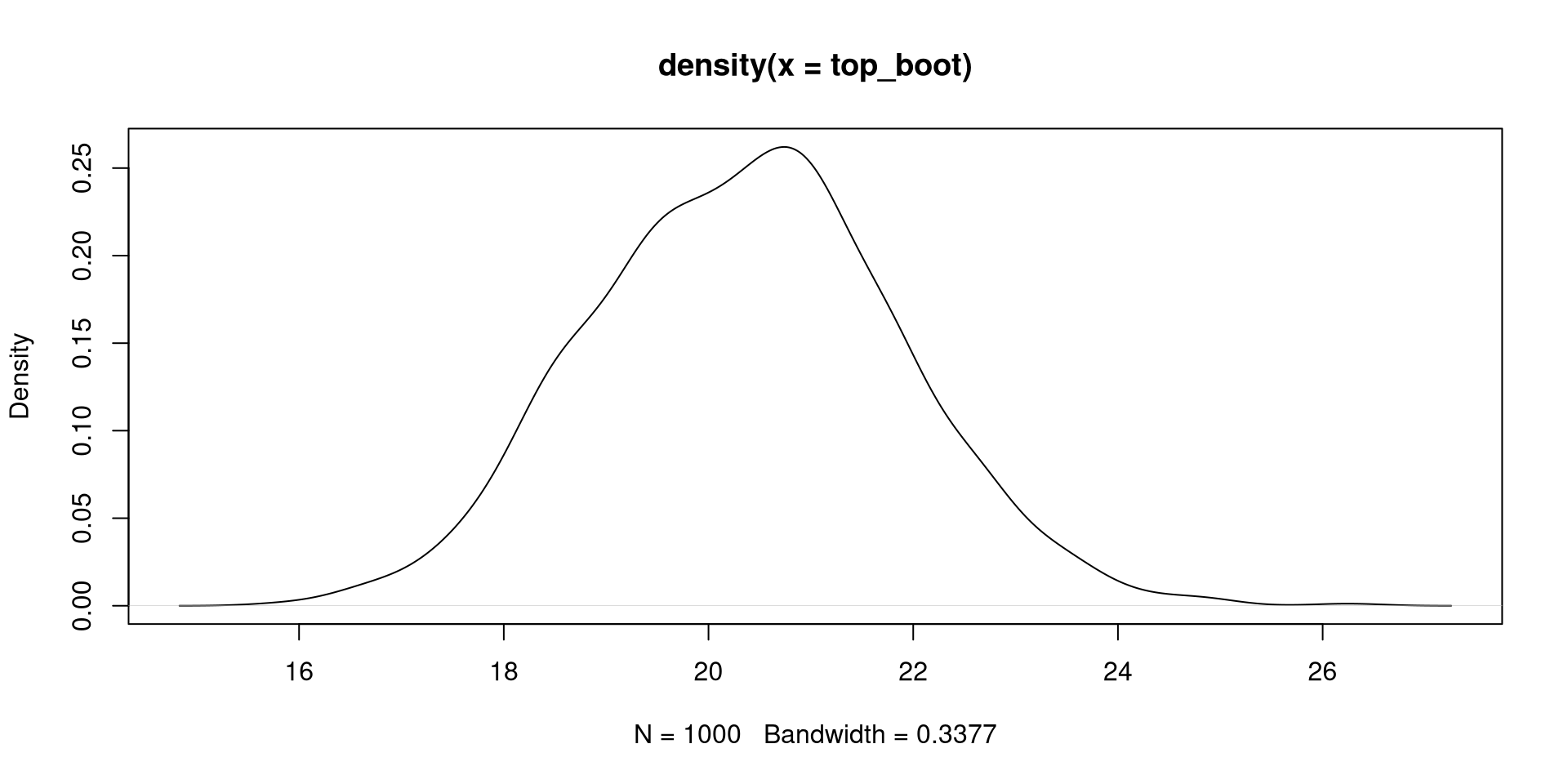
Bootsrap CI for Maximum/Minimum point
We can create a 95% CI for the time value at which we expect the maximum temperature with bootstrapping.
t element of boot_quad:
How Many Polynomials do I need?
It is my opinion that will almost never need more than a quadratic term if you are doing research in Psychology. In our case, we are dealing with a physical phenomenon (temperature), so higher polynomials may not be as far fetched.
poly() function:
The poly() Function
Be careful! It may seem that the poly() function saves us a lot of time, but it calculates orthogonal polynomials. Do not interpret the regression coefficients of regressions that use poly() functions! For more, see this link.
Comparing Polynomial Regressions
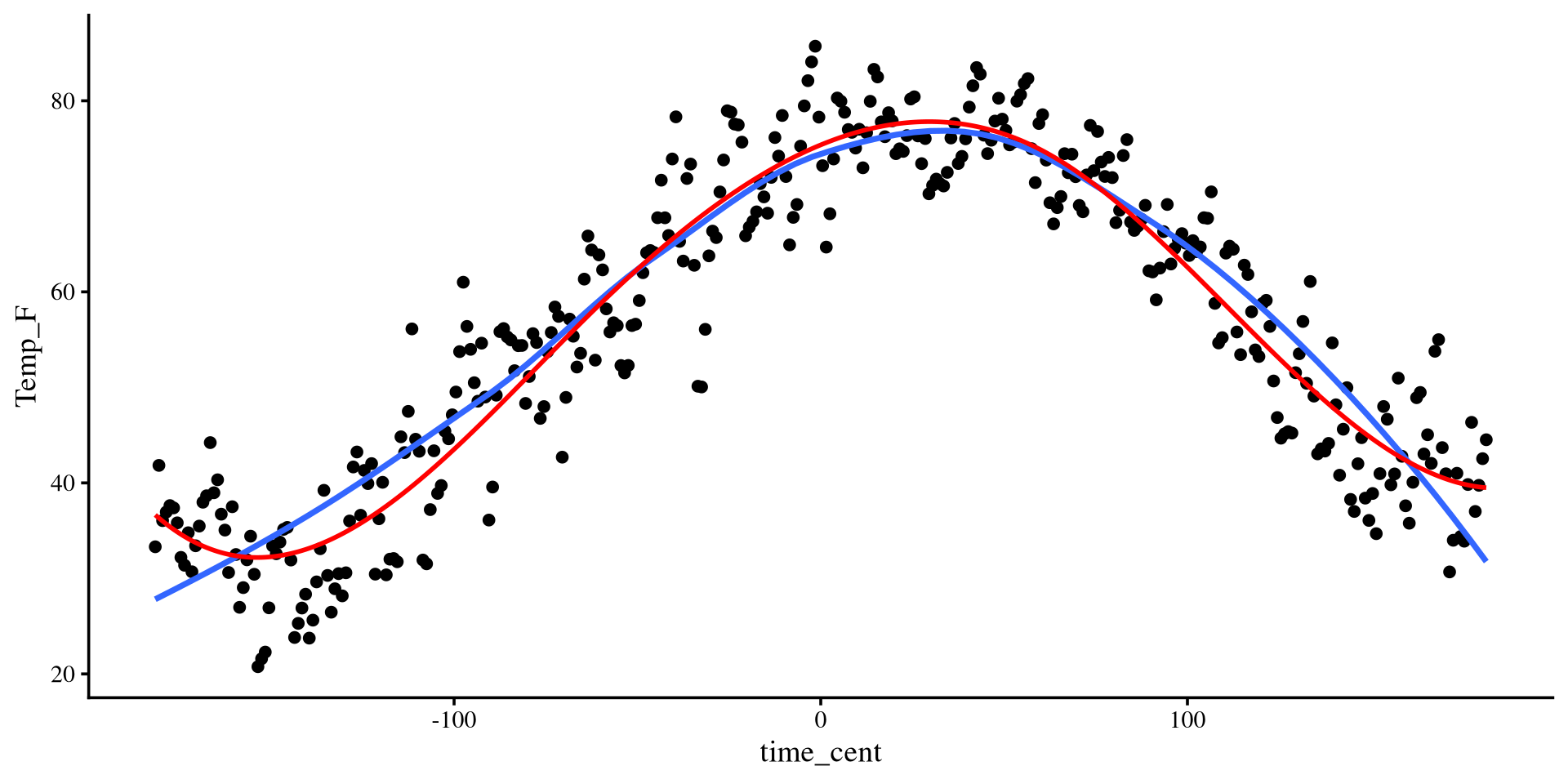
The \(\Delta R^2\) suggests that there is a significant \(R^2\) improvement up to a polynomial of the 4th degree. Here are the \(R^2\) values of the 5 regression models:
[1] 0.1233529 0.7815182 0.8341097 0.8943139 0.8950277The 5th degree polynomial offers essentially no improvement.
Analysis of Variance Table
Model 1: Temp_F ~ time
Model 2: Temp_F ~ poly(time, 2)
Model 3: Temp_F ~ poly(time, 3)
Model 4: Temp_F ~ poly(time, 4)
Model 5: Temp_F ~ poly(time, 5)
Res.Df RSS Df Sum of Sq F Pr(>F)
1 362 88352
2 361 22019 1 66332 2244.6235 <2e-16 ***
3 360 16719 1 5300 179.3593 <2e-16 ***
4 359 10651 1 6068 205.3219 <2e-16 ***
5 358 10579 1 72 2.4343 0.1196
---
Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1Let’s quickly look at what the polynomial regression lines look like.
Plotting polynomials
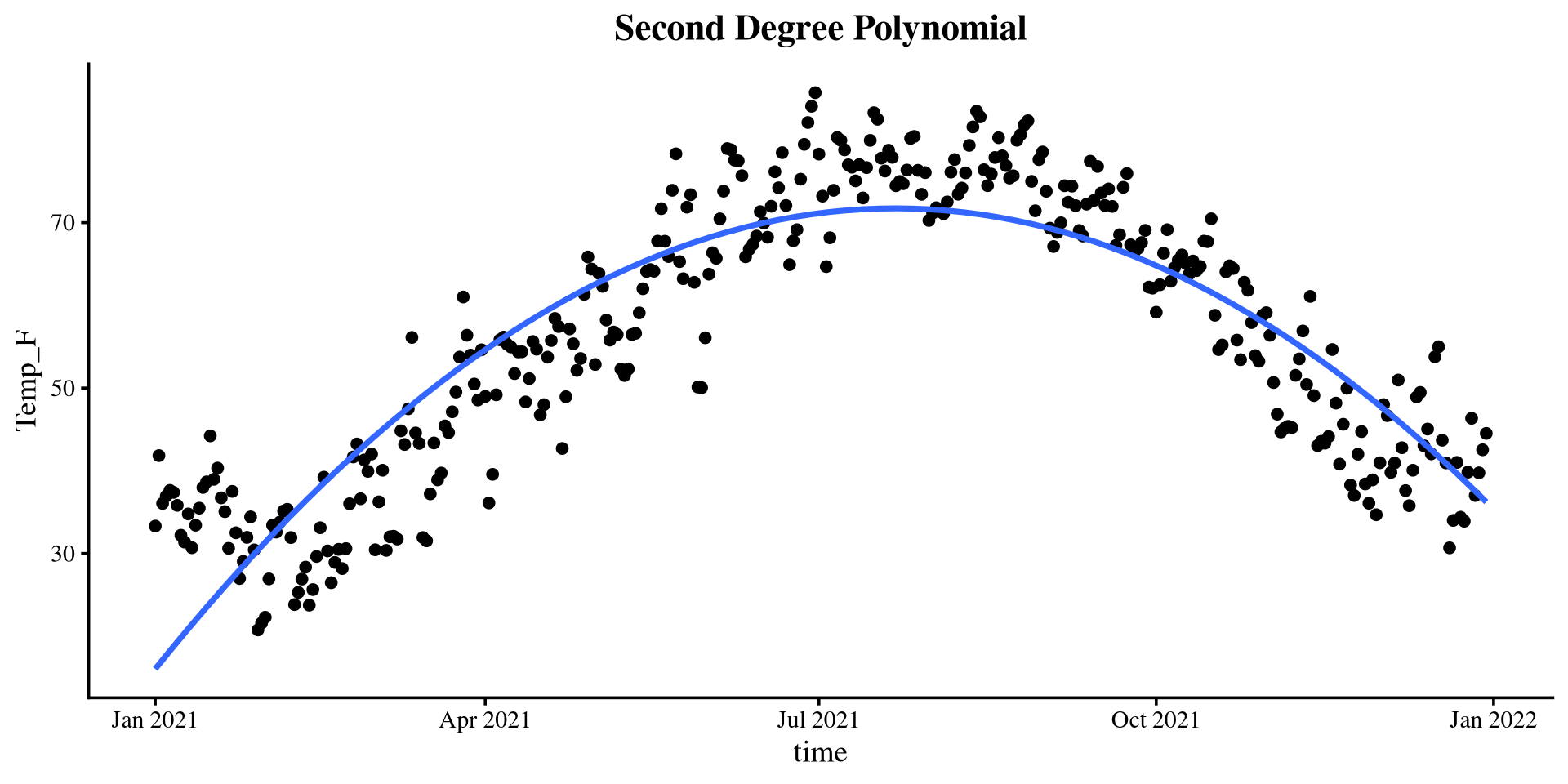
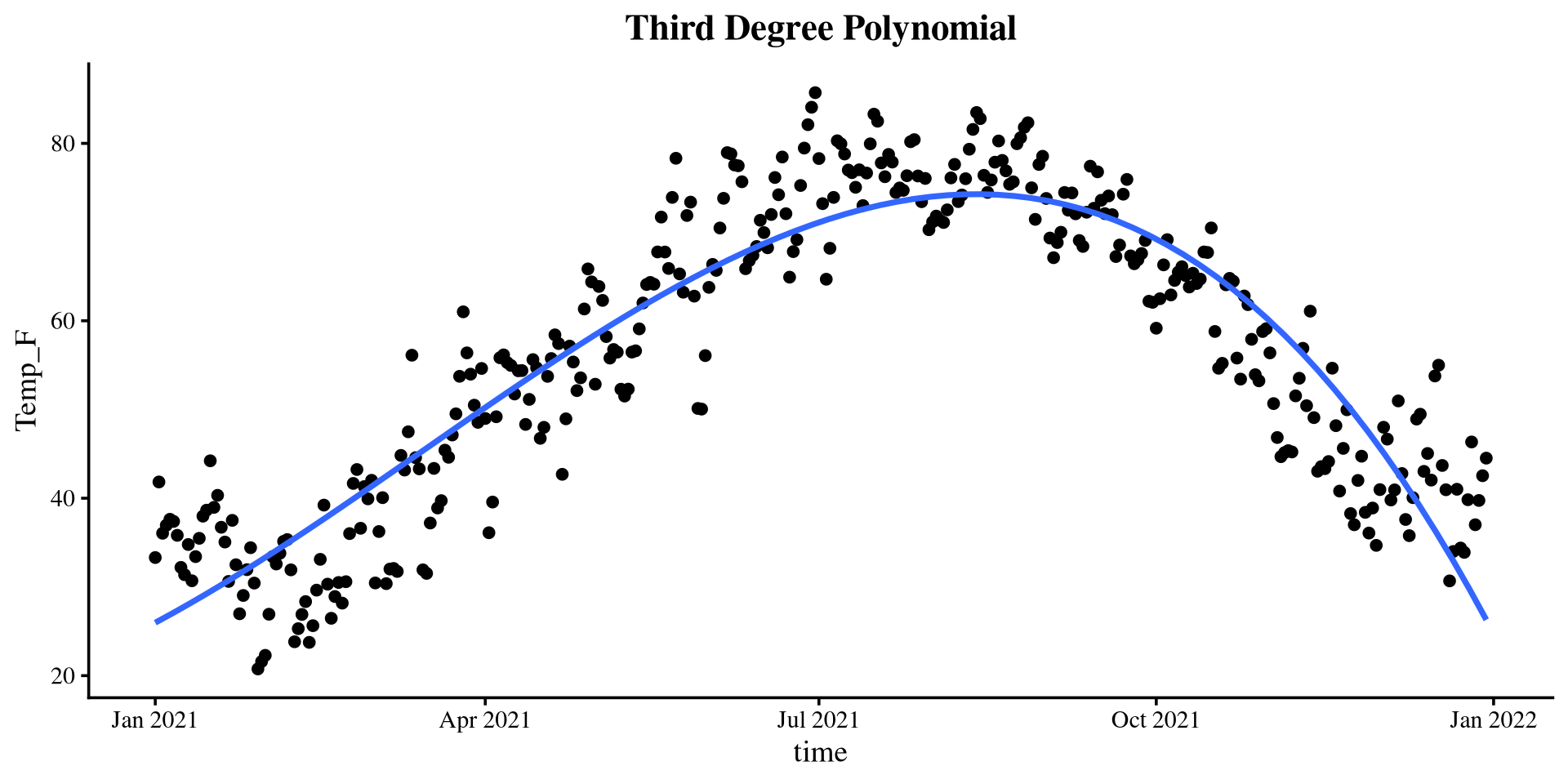
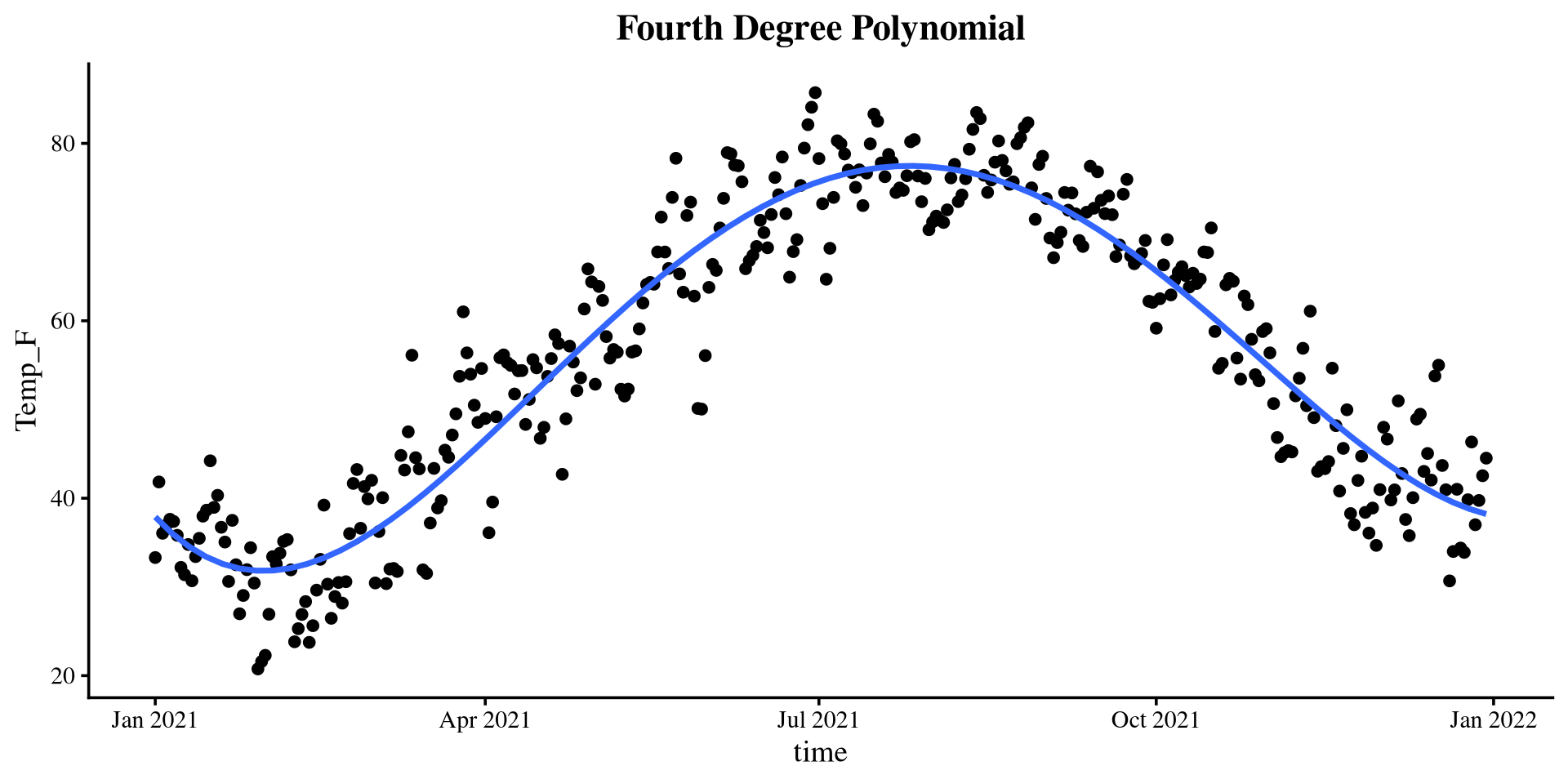
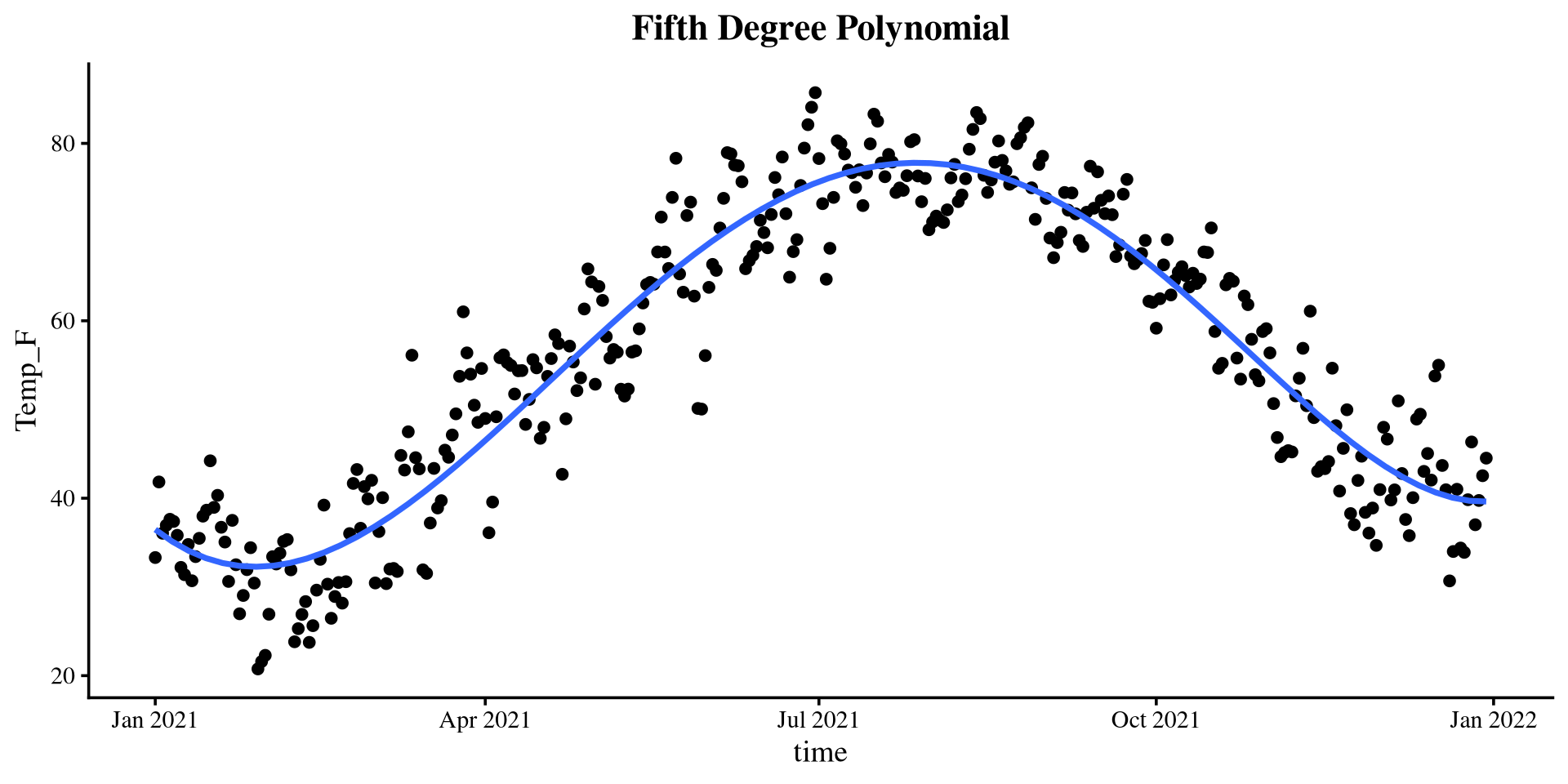
Piecewise Regression
- Why? Because we know that the top is the turning point of the quadratic line. If I need to fit two lines to the data, that is probably my best option given what I know.
Plot code
# split data with cut-point (cleaner code in 2 slides, can't be bothered to change it)
Temp_avg_2021$D1 <- ifelse(Temp_avg_2021$time_cent >= 20.34353, 0*Temp_avg_2021$time_cent, 1*Temp_avg_2021$time_cent)
Temp_avg_2021$D2 <- ifelse(Temp_avg_2021$time_cent < 20.34353, 0*Temp_avg_2021$time_cent, 1*Temp_avg_2021$time_cent)
reg_piece <- lm(Temp_F ~ D1 + D2, data = Temp_avg_2021)
intercept <- coef(reg_piece)[1]
D1 <- coef(reg_piece)[2]
D2 <- coef(reg_piece)[3]
ggplot(Temp_avg_2021,
aes(x = time, y = Temp_F)) +
geom_point() +
geom_segment(x = min(Temp_avg_2021$time_cent) + mean(Temp_avg_2021$time),
xend = 20.34353 + mean(Temp_avg_2021$time),
y = intercept + min(Temp_avg_2021$time_cent)*D1 ,
yend = intercept,
linetype = 1,
linewidth = .7,
col = "red") +
geom_segment(xend = max(Temp_avg_2021$time_cent) + mean(Temp_avg_2021$time),
x = 20.34353 + mean(Temp_avg_2021$time),
yend = intercept + max(Temp_avg_2021$time_cent)*D2 ,
y = intercept,
linetype = 1,
linewidth = .7,
col = "red") +
theme(plot.title = element_text(hjust = 0.5, face = "bold"))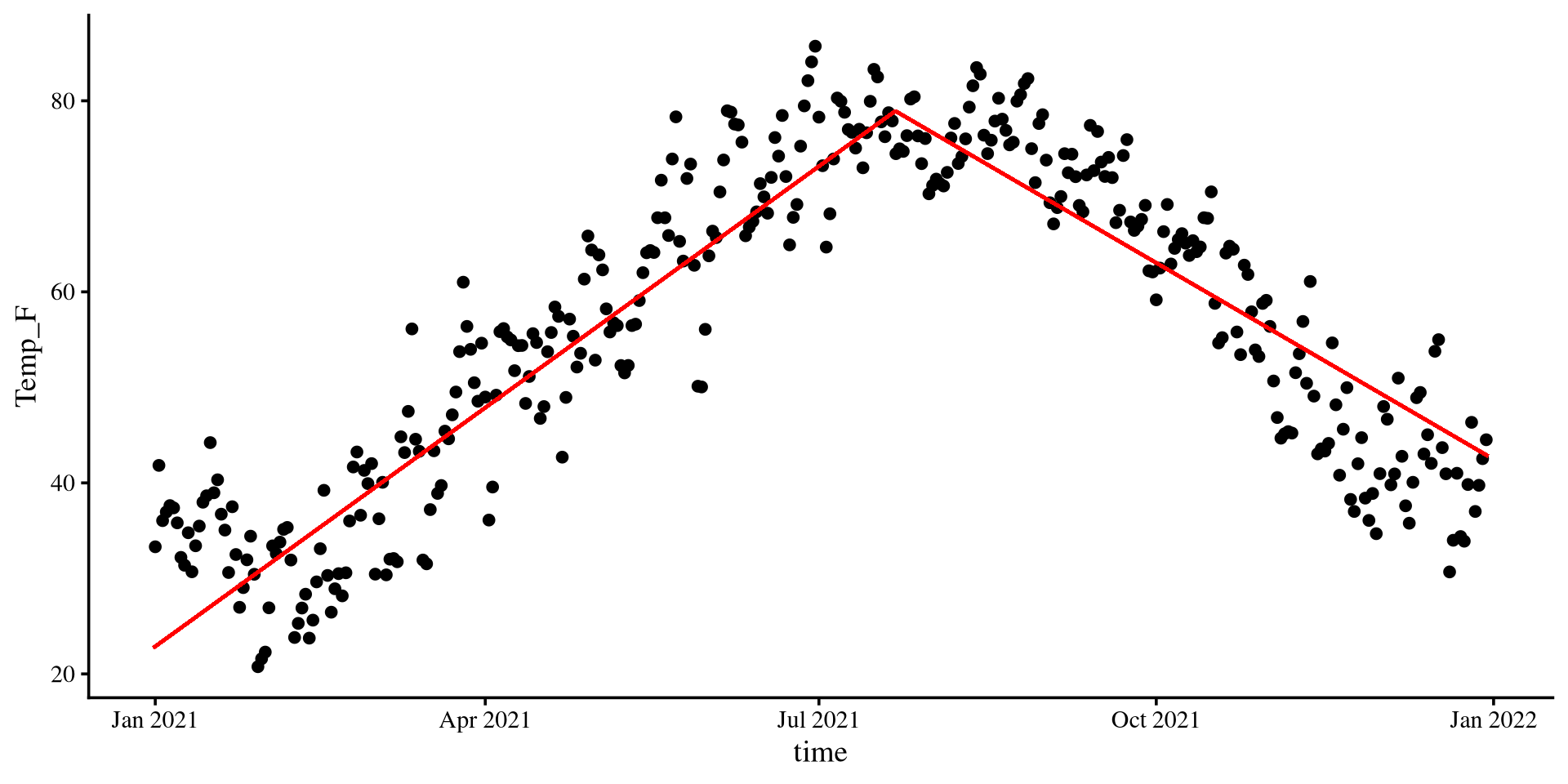
Setting up Piecewise
There are multiple ways to set up piecewise regression in R. I find the following method the most intuitive:
\[ \hat{Y} = b_0 + b_1\times D_1 + b_2 \times D_2 \]
We need to create \(D_1\) and \(D_2\). The trick is to get the original predictor, time_cent in our case, and split its value across the segments.
So, if we choose our cut-point to be around the top or the quadratic curve, roughly \(20.34\):
- \(D_1:\) Make all values after \(20.34\) become \(0\).
- \(D_2:\) Make all values before \(20.34\) become \(0\).
We will use the ifelse() function to make our life easier.
Creating \(D_1\) and \(D_2\)
- Creating \(D_1\):
- Creating \(D_2\):
temp_cent variable has been split between \(D_1\) and \(D_2\).
Running The Piecewise Regression
Now we can simply run our piecewise regression by using \(D_1\) and \(D_2\) as predictors:
Call:
lm(formula = Temp_F ~ D1 + D2, data = Temp_avg_2021)
Residuals:
Min 1Q Median 3Q Max
-18.8270 -5.4317 -0.1449 5.5464 18.7796
Coefficients:
Estimate Std. Error t value Pr(>|t|)
(Intercept) 78.933629 0.728574 108.34 <2e-16 ***
D1 0.309589 0.007800 39.69 <2e-16 ***
D2 -0.199029 0.007816 -25.46 <2e-16 ***
---
Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
Residual standard error: 7.196 on 361 degrees of freedom
Multiple R-squared: 0.8145, Adjusted R-squared: 0.8135
F-statistic: 792.6 on 2 and 361 DF, p-value: < 2.2e-163D visualization? Out of Curiosity…
The 3D visualization of a piecewise regression is actually kinda funny:
Spline Regression
Spline regression is quite similar to loess regression. Spline regression is a piecewise regression with many segments, and is very similar to loess regression.
Call:
lm(formula = Temp_F ~ splines2::bsp(Temp_avg_2021$time_cent,
df = 5), data = Temp_avg_2021)
Residuals:
Min 1Q Median 3Q Max
-17.9151 -3.8278 0.1797 3.8033 16.5899
Coefficients:
Estimate Std. Error t value
(Intercept) 36.645 1.577 23.230
splines2::bsp(Temp_avg_2021$time_cent, df = 5)1 -13.622 2.989 -4.557
splines2::bsp(Temp_avg_2021$time_cent, df = 5)2 27.560 1.981 13.912
splines2::bsp(Temp_avg_2021$time_cent, df = 5)3 55.720 2.728 20.423
splines2::bsp(Temp_avg_2021$time_cent, df = 5)4 4.128 2.213 1.865
splines2::bsp(Temp_avg_2021$time_cent, df = 5)5 2.851 2.301 1.239
Pr(>|t|)
(Intercept) < 2e-16 ***
splines2::bsp(Temp_avg_2021$time_cent, df = 5)1 7.13e-06 ***
splines2::bsp(Temp_avg_2021$time_cent, df = 5)2 < 2e-16 ***
splines2::bsp(Temp_avg_2021$time_cent, df = 5)3 < 2e-16 ***
splines2::bsp(Temp_avg_2021$time_cent, df = 5)4 0.063 .
splines2::bsp(Temp_avg_2021$time_cent, df = 5)5 0.216
---
Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
Residual standard error: 5.423 on 358 degrees of freedom
Multiple R-squared: 0.8955, Adjusted R-squared: 0.8941
F-statistic: 613.8 on 5 and 358 DF, p-value: < 2.2e-16We can use the bsp() function form the splines2 package to define our spline regression. the df = arguments sets the number of independent segments. here I choose 5 segments.
The coefficients are not that helpful. Let’s look at the visualization.
Visulizing Spline Regression
The visualization for spline regression requires you calculate the predictions from the regression model (unless I decide to make a function at some point):
slpine2 package uses 3rd degree polynomial segments. (see help(bsp))
References
PSYC 7804 - Lab 8: Quadratic regression and non-linear alternatives