Lab 4: Introduction To Two-Predictor Regression
PSYC 7804 - Regression with Lab
Spring 2025
Today’s Packages and Data 🤗
The GGally package (Schloerke et al., 2024) builds upon ggplot2 and includes many functions for creating complex plots.
The plotly package (Sievert et al., 2024) is a Python and R package used to create interactive visualizations with JavaScript elements.
'data.frame': 140 obs. of 9 variables:
$ Country_name : chr "Finland" "Denmark" ...
$ Region : chr "Western Europe" "Western Europe" ...
$ Happiness_score : num 7.74 7.58 ...
$ Log_GDP : num 1.84 1.91 ...
$ Social_support : num 1.57 1.52 ...
$ Healthy_life_expectancy: num 0.695 0.699 0.718 0.724 0.74 ...
$ Freedom : num 0.859 0.823 0.819 0.838 0.641 ...
$ Generosity : num 0.142 0.204 0.258 0.221 0.153 ...
$ Corruption : num 0.454 0.452 0.818 0.476 0.807 ...Variables of Interest
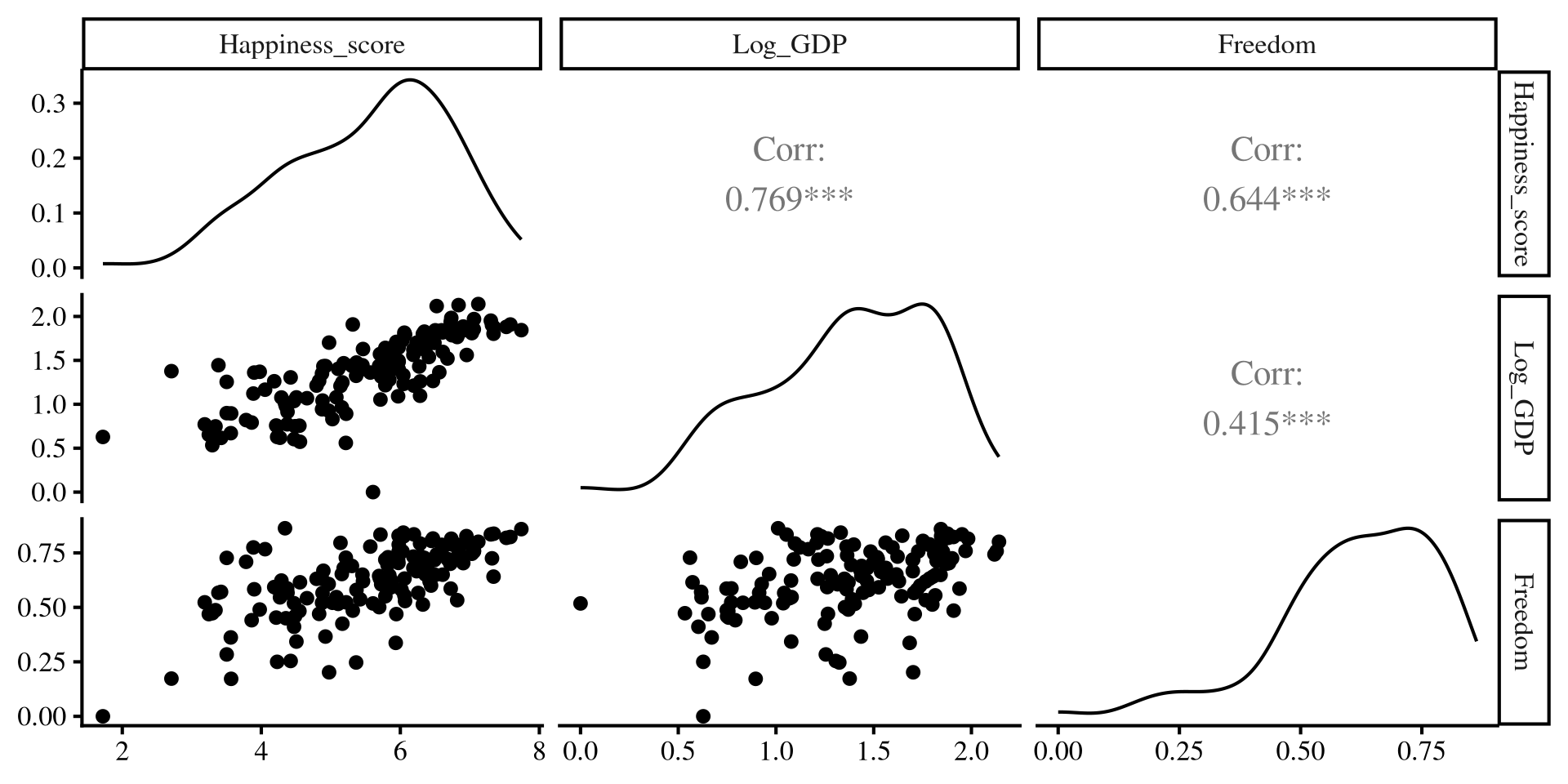
We eventually want to look at how Log_GDP and Freedom impact Happiness_score across the countries in our data. Let’s first explore the variables.
- The correlation matrix for the variables:
Happiness_score Log_GDP Freedom
Happiness_score 1.0000000 0.7685037 0.6444511
Log_GDP 0.7685037 1.0000000 0.4148856
Freedom 0.6444511 0.4148856 1.0000000- Some descriptive statistics:
vars n mean sd median trimmed mad min max range skew
Happiness_score 1 140 5.53 1.18 5.80 5.60 1.21 1.72 7.74 6.02 -0.52
Log_GDP 2 140 1.38 0.43 1.43 1.40 0.50 0.00 2.14 2.14 -0.50
Freedom 3 140 0.62 0.16 0.64 0.64 0.15 0.00 0.86 0.86 -1.00
kurtosis se
Happiness_score -0.29 0.10
Log_GDP -0.42 0.04
Freedom 1.16 0.01Helpful Visualization
The ggpairs() function from the GGally package creates a visualizations that provides a lot of information in one go!
Individual Regressions
-
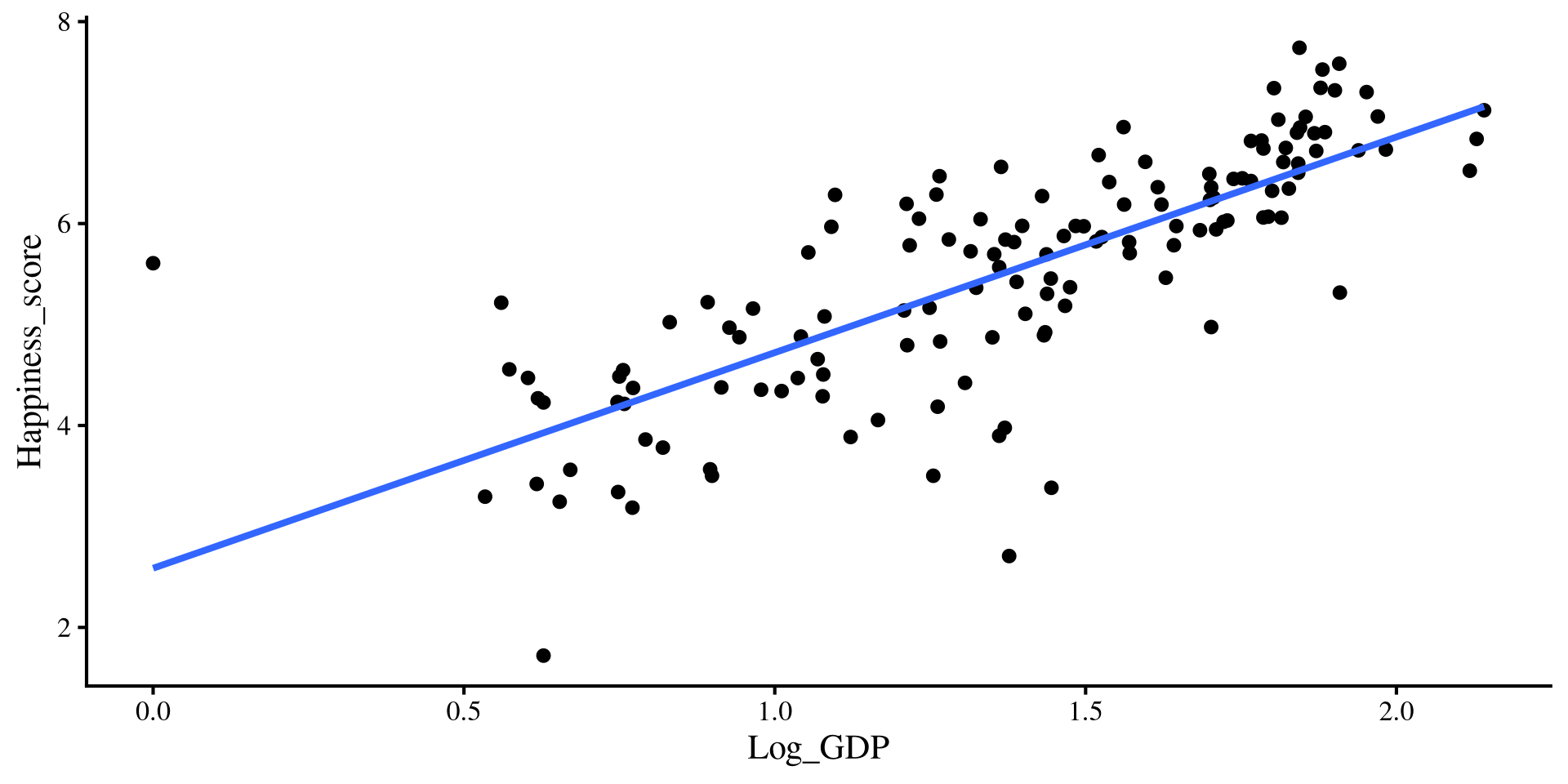
Log_GDPonly regression
Call:
lm(formula = Happiness_score ~ Log_GDP, data = reg_vars)
Residuals:
Min 1Q Median 3Q Max
-2.82003 -0.37094 0.05679 0.40998 3.02053
Coefficients:
Estimate Std. Error t value Pr(>|t|)
(Intercept) 2.5865 0.2183 11.85 <2e-16 ***
Log_GDP 2.1355 0.1514 14.11 <2e-16 ***
---
Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
Residual standard error: 0.7585 on 138 degrees of freedom
Multiple R-squared: 0.5906, Adjusted R-squared: 0.5876
F-statistic: 199.1 on 1 and 138 DF, p-value: < 2.2e-16-
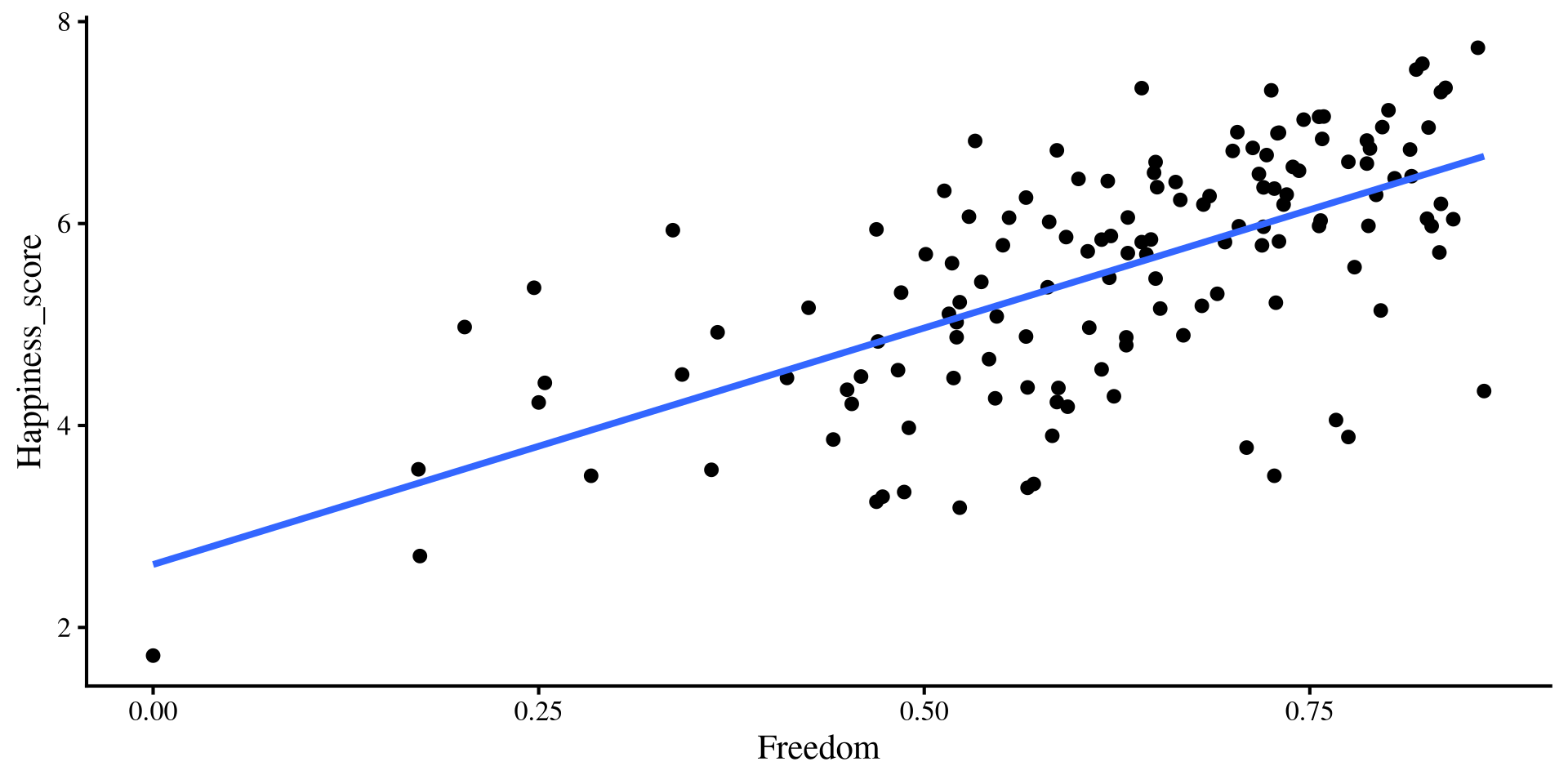
Freedomonly regression
Call:
lm(formula = Happiness_score ~ Freedom, data = reg_vars)
Residuals:
Min 1Q Median 3Q Max
-2.5273 -0.5317 0.1337 0.6656 1.7318
Coefficients:
Estimate Std. Error t value Pr(>|t|)
(Intercept) 2.6234 0.3035 8.644 1.22e-14 ***
Freedom 4.6849 0.4732 9.901 < 2e-16 ***
---
Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
Residual standard error: 0.9065 on 138 degrees of freedom
Multiple R-squared: 0.4153, Adjusted R-squared: 0.4111
F-statistic: 98.03 on 1 and 138 DF, p-value: < 2.2e-16Regression With Both Predictors
Call:
lm(formula = Happiness_score ~ Log_GDP + Freedom, data = reg_vars)
Residuals:
Min 1Q Median 3Q Max
-2.10592 -0.23908 0.09905 0.34516 2.68875
Coefficients:
Estimate Std. Error t value Pr(>|t|)
(Intercept) 1.4372 0.2327 6.175 7.05e-09 ***
Log_GDP 1.6821 0.1384 12.154 < 2e-16 ***
Freedom 2.8592 0.3621 7.897 8.28e-13 ***
---
Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
Residual standard error: 0.6311 on 137 degrees of freedom
Multiple R-squared: 0.7187, Adjusted R-squared: 0.7146
F-statistic: 175 on 2 and 137 DF, p-value: < 2.2e-16As you can see, the regression coefficients of the two predictors change when both variables are included 🧐
Let’s visualize what is happening!
Individual Regression plots
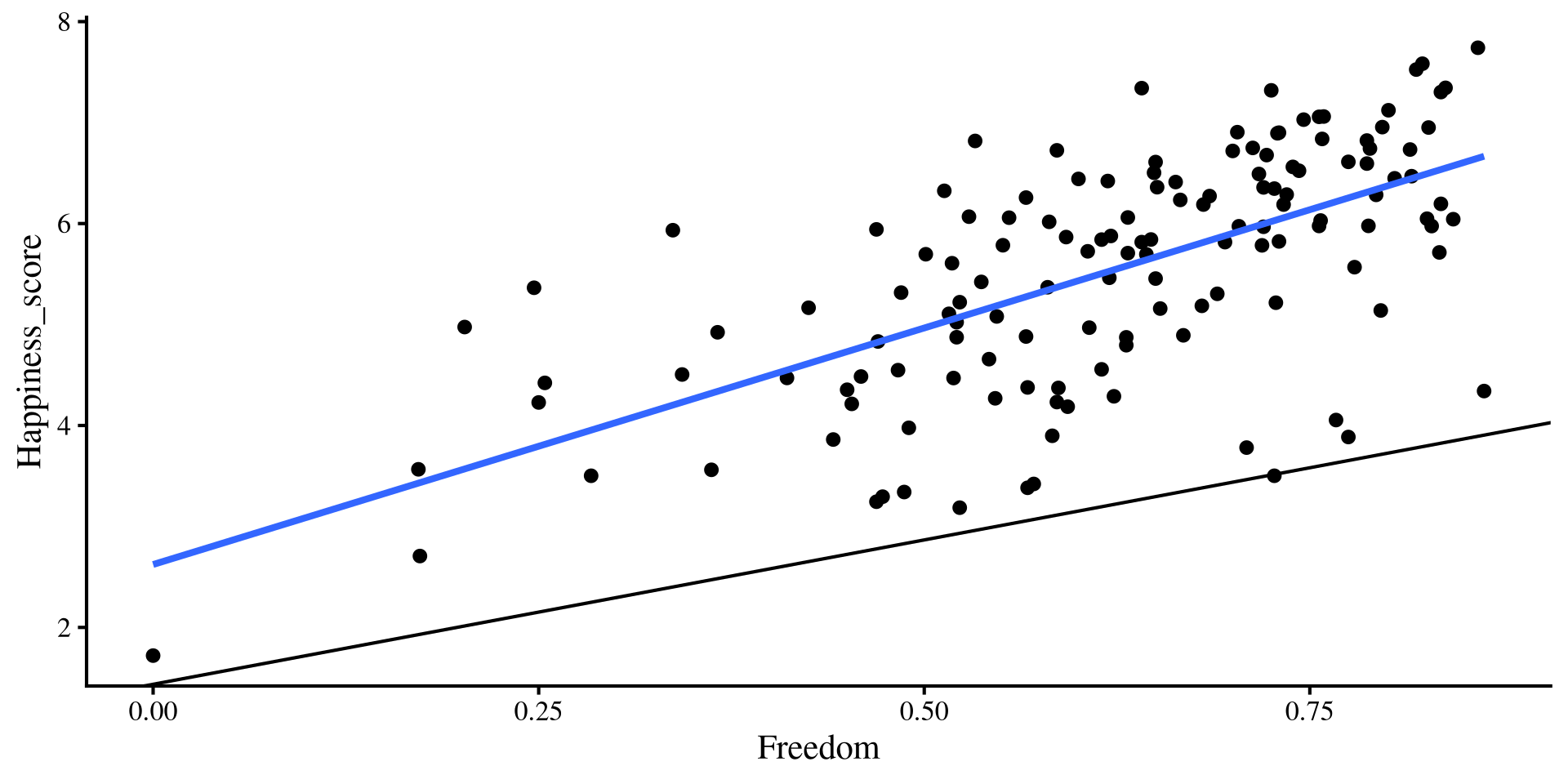
These are the equivalent plots to the individual regressions:
Adding Slopes from multiple regression?
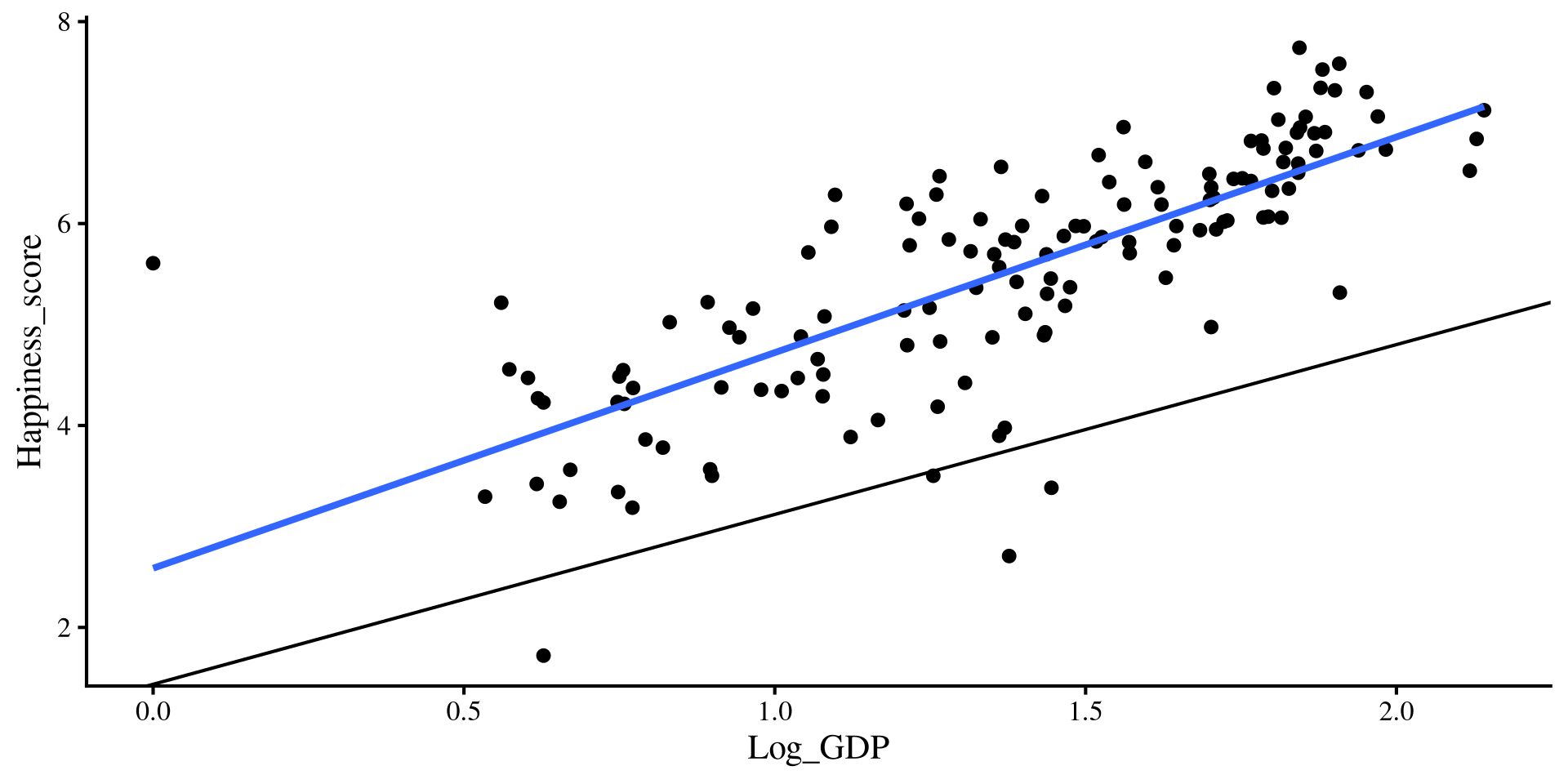
Let’s add the regression lines estimated from the regression with both predictors…
Hold up, the lines from the multiple regression results seem way off !?
💡 Maybe we need a change of perspective!
Looking Inside the Box
Now that we are dealing with 3 variables (Log_GDP, Freedom, Happiness_score), our data points are in a 3D box, not a 2D plot.
I made a function for making interactive 3D plots easily
The 2D plots from the previous slide are just the sides of the box 🤓
The Regression Plane
Remember that in linear regression with 1 predictor we are looking for the line that on average is closest to all point. When we have 2 predictors we are looking for the plane that is closest to all the points!
Question?
If this is the representation of regression with 2 predictors, what happens once we have 3 or more predictors? Can we visualize that?
Interpretation of Results
Now that we have a sense of what we are looking at, let’s interpret the output:
Call:
lm(formula = Happiness_score ~ Log_GDP + Freedom, data = reg_vars)
Residuals:
Min 1Q Median 3Q Max
-2.10592 -0.23908 0.09905 0.34516 2.68875
Coefficients:
Estimate Std. Error t value Pr(>|t|)
(Intercept) 1.4372 0.2327 6.175 7.05e-09 ***
Log_GDP 1.6821 0.1384 12.154 < 2e-16 ***
Freedom 2.8592 0.3621 7.897 8.28e-13 ***
---
Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
Residual standard error: 0.6311 on 137 degrees of freedom
Multiple R-squared: 0.7187, Adjusted R-squared: 0.7146
F-statistic: 175 on 2 and 137 DF, p-value: < 2.2e-16-
\(b_0 = 1.44\): the expected
happpiness_scorewhenGDPandFreedomare at 0. -
\(b_1 = 1.68\): the expected increase in
happpiness_scoreper each 1-unit increase inLog_GDPwhen accounting forFreedom. -
\(b_2 = 2.86\): the expected increase in
happpiness_scoreper each 1-unit increase inFreedomwhen accounting forGDP.
\(\widehat{\mathrm{Happiness}} = 1.44 + 1.68\times \mathrm{Log\_GDP} + 2.86\times \mathrm{Freedom}\)
\(b_1\) and \(b_2\) are partial regression coefficients. We will come back to this in Lab 5, so stay tuned.
Back to our friend R 2
You can think of variance as how unpredictable something is. Higher variance means more unpredictability. At the same time, no variance means perfect predictability!
For example, if you generate data for a variable that has 0 variance, you always get the mean
Here, if we predict 2 we are always right, because the variable has no variance and is perfectly predictable.
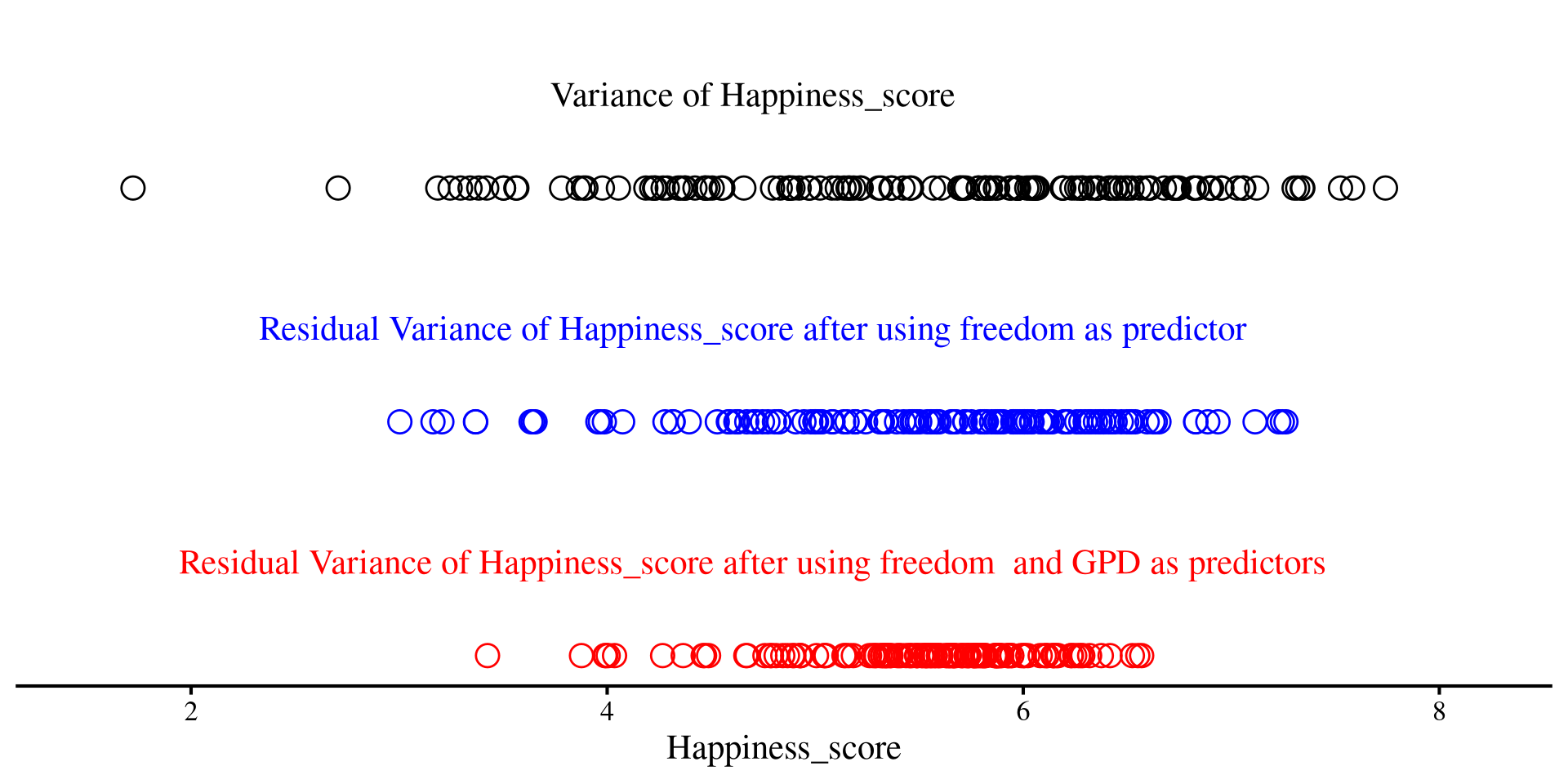
Let’s return to our 1D scatterplot, and let’s visualize the variance of the Happiness_score
Code
ggplot(reg_vars,
aes(x = Happiness_score, y = 0)) +
geom_point(shape = 1,
size=6.5) +
annotate("text", x = 4.7, y = .02,
label = "Variance of Happiness_score") +
xlim(1.5, 8.2) +
ylim(-.1, .03) +
theme(axis.title.y=element_blank(),
axis.text.y=element_blank(),
axis.ticks.y=element_blank(),
axis.line.y = element_blank())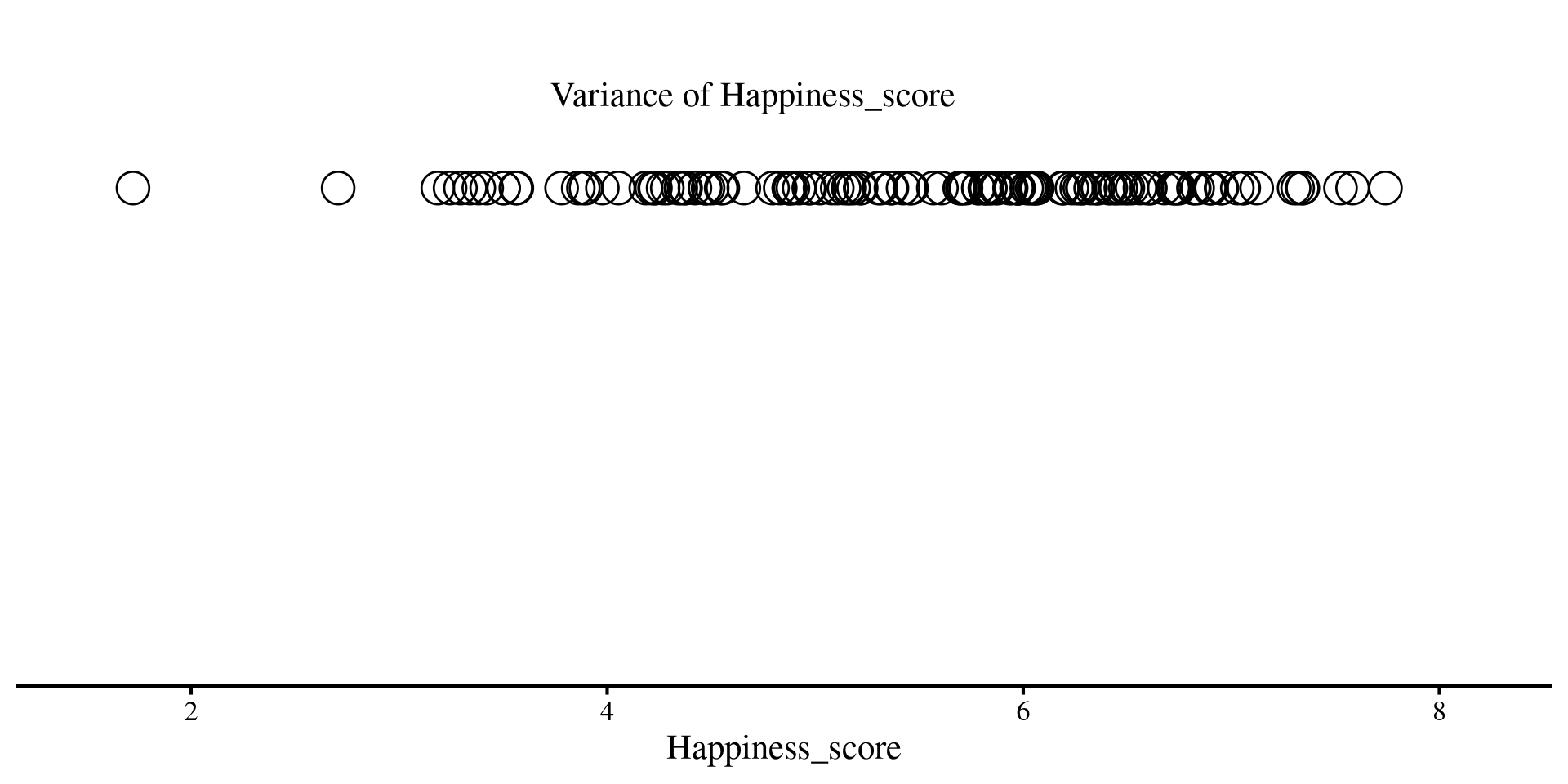
Variance of Residuals after regression
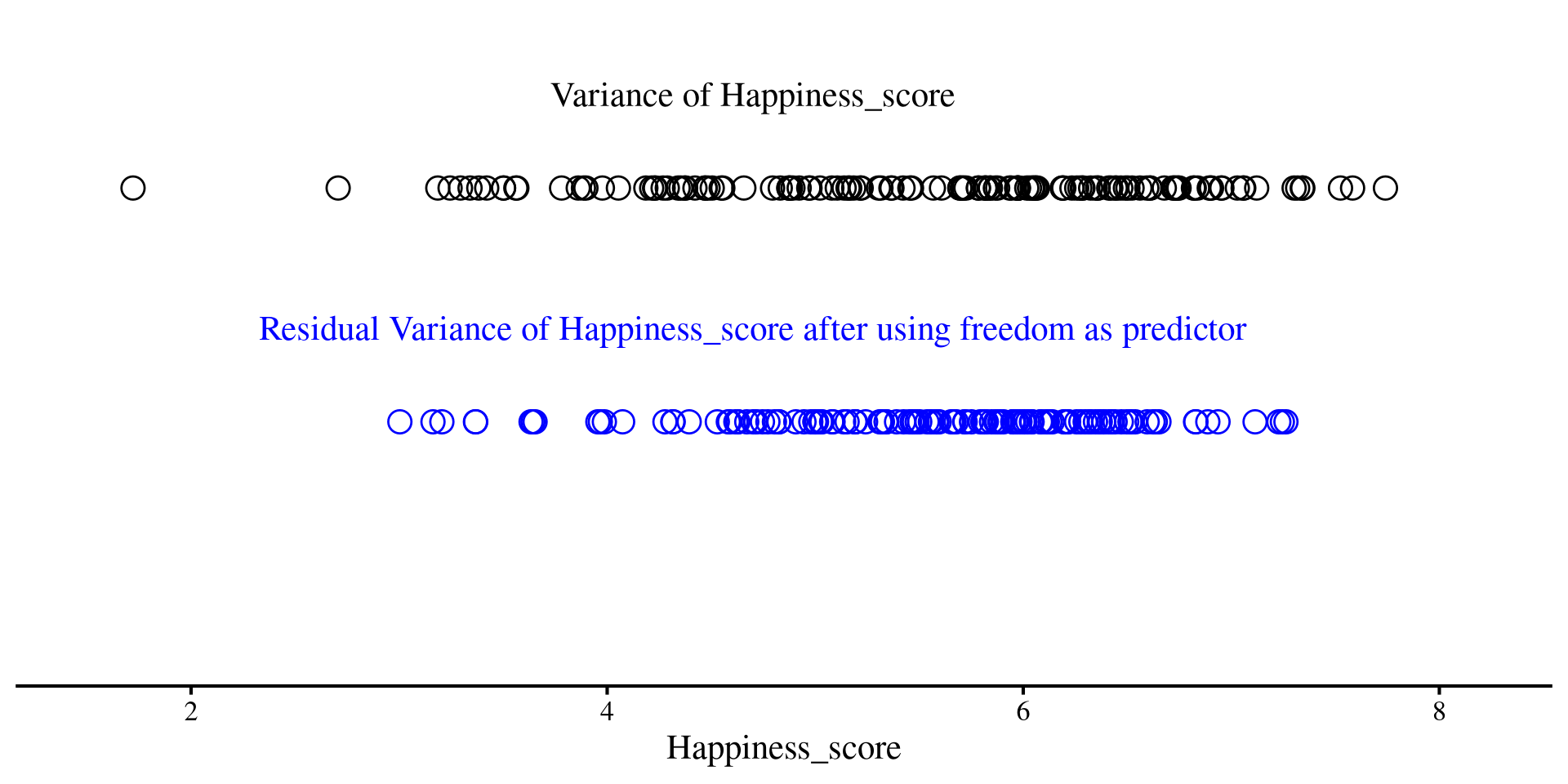
Although this may not sound very intuitive, you should think of residuals as a new version of your \(Y\) variable, but with some unpredictability taken out thanks to our regression line. Less unpredictability means lower variance
On the right, we have the residuals of Happiness_score after using freedom as predictor
Code
ggplot(reg_vars,
aes(x = Happiness_score, y = 0)) +
geom_point(shape = 1,
size=4.5) +
annotate("text", x = 4.7, y = .02,
label = "Variance of Happiness_score") +
geom_point(aes(x = residuals(reg_free) +
# add mean to have residuals on same scale as
# previous graph (variance remains the same)
mean(reg_vars$Happiness_score),
y = -.05),
shape = 1,
size = 4.5, color = "blue") +
annotate("text", x = 4.7, y = -0.03,
label = "Residual Variance of Happiness_score after using freedom as predictor", color = "blue") +
xlim(1.5, 8.2) +
ylim(-.1, .03) +
theme(axis.title.y=element_blank(),
axis.text.y=element_blank(),
axis.ticks.y=element_blank(),
axis.line.y = element_blank())Much less spread out, right?
Variance of Residuals after Regression
Although this may not sound very intuitive, you should think of residuals as a new version of your \(Y\) variable, but with some unpredictability taken out thanks to our regression line. Less unpredictability means lower variance
Now, after we add both freedom and GDP as predictors, we see that the dots are even less spread out
Code
ggplot(reg_vars,
aes(x = Happiness_score, y = 0)) +
geom_point(shape = 1,
size=4.5)+
annotate("text", x = 4.7, y = .02,
label = "Variance of Happiness_score") +
geom_point(aes(x = residuals(reg_free) +
# add mean to have residuals on same scale as
# original variable (variance remains the same)
mean(reg_vars$Happiness_score),
y = -.05),
shape = 1,
size = 4.5, color = "blue") +
annotate("text", x = 4.7, y = -0.03,
label = "Residual Variance of Happiness_score after using freedom as predictor", color = "blue") +
geom_point(aes(x = residuals(reg_full) +
# add mean to have residuals on same scale as
# original variable (variance remains the same)
mean(reg_vars$Happiness_score),
y = -.1),
shape = 1,
size = 4.5, color = "red") +
annotate("text", x = 4.7, y = -0.08,
label = "Residual Variance of Happiness_score after using freedom and GPD as predictors", color = "red") +
xlim(1.5, 8.2) +
ylim(-.1, .03) +
theme(axis.title.y=element_blank(),
axis.text.y=element_blank(),
axis.ticks.y=element_blank(),
axis.line.y = element_blank())The red dots are even less spread out than the blue ones!
Variance Explained
As we saw previously, the more variables we added, the less spread out our residuals were. A part of the variance of the original variable was explained by our predictors.
The variances of the three lines of dots were respectively:
-
Variance of
happiness_scoreafter usingfreedomas a predictor was
We can easily calculate \(R^2\) by hand
Significance For R 2
As mentioned in Lab 3, significance testing targets the sampling distribution of a statistic. For regression coefficients, the sampling distribution is a \(t\)-distribution. For \(R^2\), we use an \(F-\)distribution.
Now, why we use an \(F-\)distribution takes a few steps to explain, so I will go on a bit of a tangent.
- As we just saw, \(R^2\) tells you the percentage of variance (i.e., unpredictability) that was taken out of \(Y\) by using all the predictors in a regression. How do we tests that the \(R^2\) is not \(0\) (null hypothesis)?
-
Some smart people have figured out that we can do a “transformation” of \(R^2\),
\[F = \frac{\mathrm{df_{residual}}\times R^2}{k(1 - R^2)},\] Where \(k\) is the number of predictors, and if \(N\) is the sample size, \(\mathrm{df_{residual}} = N - k - 1\). - And it also turns out that if \(R^2\) is truly \(0\) and \(H_0\) is true, we would expect to see \(F = \frac{\mathrm{df_{residual}}}{\mathrm{df_{residual}} - 2}\). this is usually around \(F = 1\).
Why \(F\) and not \(R^2\) Directly?
Why do we got thorough all this trouble to calculate \(F\) and confuse students taking regression courses?
The formula for \(F\) on the previous slide accounts for a very important fact that we will return to in Lab 6: \(R^2\) always increases when you add additional predictors, even if the new predictors are completely unrelated to \(Y\).
The sampling distribution for an \(F\)-statistic calculated when \(H_0\) is true and \(R^2 = 0\) is, as the name suggests, an \(F\)-distribution with \(\mathrm{df}_1 = k\), and \(\mathrm{df}_2 = \mathrm{df_{residual}}\).
For significance testing, the further an \(F\)-value is from 1, the smaller the \(p\)-value will be.
ANOVA and all the \(F\)s
If you remember looking at ANOVA results, you should remember seeing a whole bunch of \(F\)-statistics. As we will see, ANOVA is a regression, but it focuses on variance explained (\(R^2\)!) by its predictors rather than regression coefficients. That is why all you will see from ANOVA are \(F\)-statistics.
P-value from the \(F\)-distribution
We want to calculate how likely it is to get our \(F\)-value from an \(F\)-distribution with \(\mathrm{df}_1 = k = 2\), and \(\mathrm{df}_2 = \mathrm{df_{residual}} = 137\).
If we look at our full regression, our \(F\)-value was \(\frac{\mathrm{df_{residual}}\times R^2}{k(1 - R^2)} = \frac{137\times .63}{2(1 - .63)} \approx 175\)
Looking at the regression Ouput
If we look at the regression output again, we will see that \(\mathrm{df}_1 = 2\), \(\mathrm{df_{residual}} = 137\), \(F = 175\), and \(p = 1.86\times10^{-38}\) are in the line below the \(R^2\) value.
Call:
lm(formula = Happiness_score ~ Log_GDP + Freedom, data = reg_vars)
Residuals:
Min 1Q Median 3Q Max
-2.10592 -0.23908 0.09905 0.34516 2.68875
Coefficients:
Estimate Std. Error t value Pr(>|t|)
(Intercept) 1.4372 0.2327 6.175 7.05e-09 ***
Log_GDP 1.6821 0.1384 12.154 < 2e-16 ***
Freedom 2.8592 0.3621 7.897 8.28e-13 ***
---
Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
Residual standard error: 0.6311 on 137 degrees of freedom
Multiple R-squared: 0.7187, Adjusted R-squared: 0.7146
F-statistic: 175 on 2 and 137 DF, p-value: < 2.2e-16In APA style you would report this at the end of the regression results as:
…the model explained a significant porportion of variance in country happiness, \(R^2 = .72\), \(F\)(2, 137) = 175, \(p < .001\).
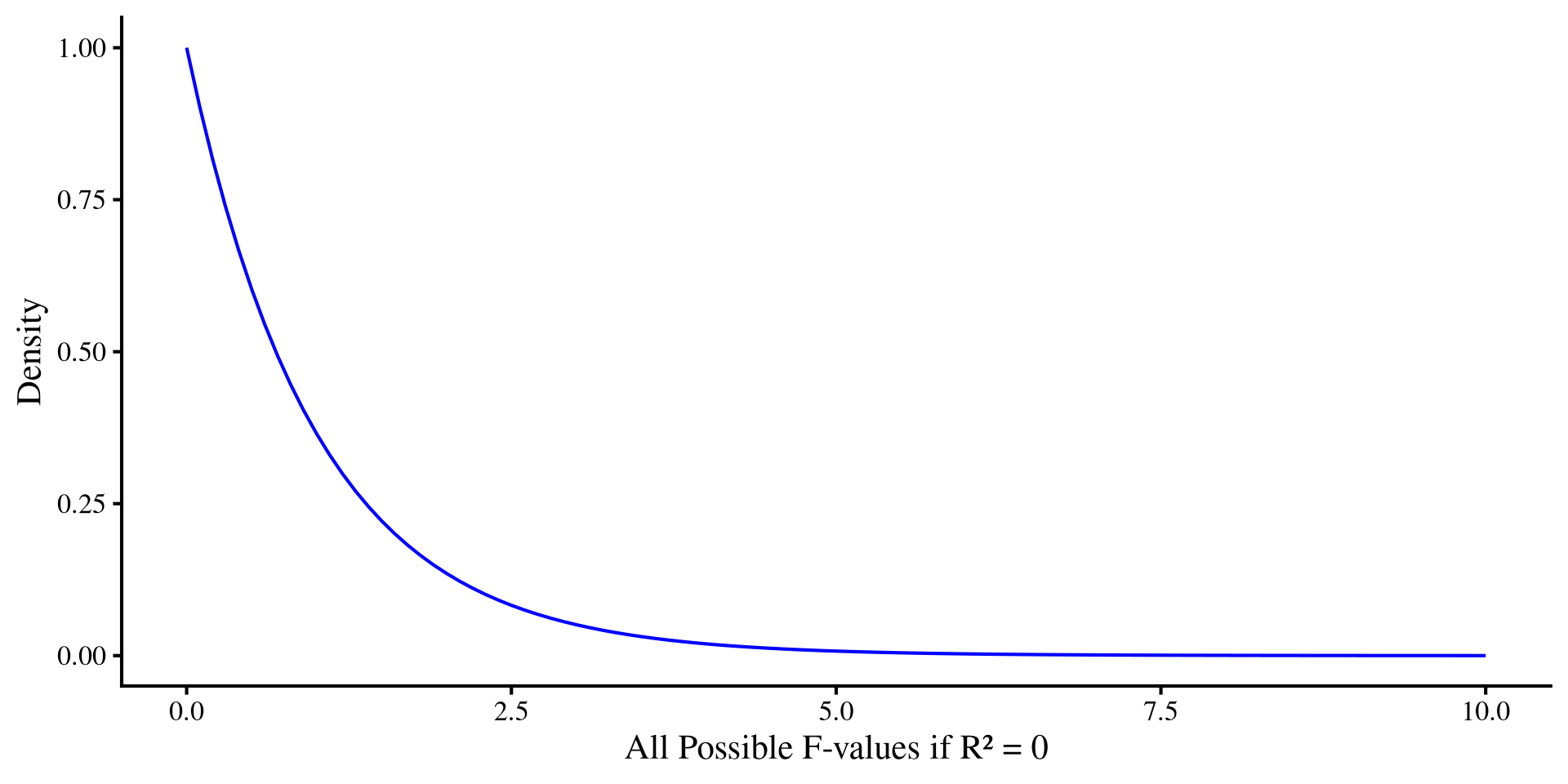
What does the \(F\)-distribution look like?
We can see what the \(F\)-distribution looks like when we change degrees of freedom
\(\mathrm{df}_1\), representing number of variables, is much more influential on the shape compared to \(\mathrm{df}_2\), which represents the sample size.
Simulating Experiments where \(H_0\) is true
sample_size <- 300
# object where we save F-statistic
F_statistics <- c()
for(i in 1:2000){
X <- rnorm(sample_size, mean = 0, sd = 1)
# the regression coefficient of X is exactly 0
Y <- rnorm(sample_size, mean = 0*X, sd = 1)
reg_YX <- lm(Y ~ X)
# store F-statistic at every iteration
F_statistics[i] <- as.numeric(summary(reg_YX)$fstatistic[1])
}
ggplot() +
geom_density(aes(x = F_statistics)) +
scale_y_continuous(expand = c(0, 0))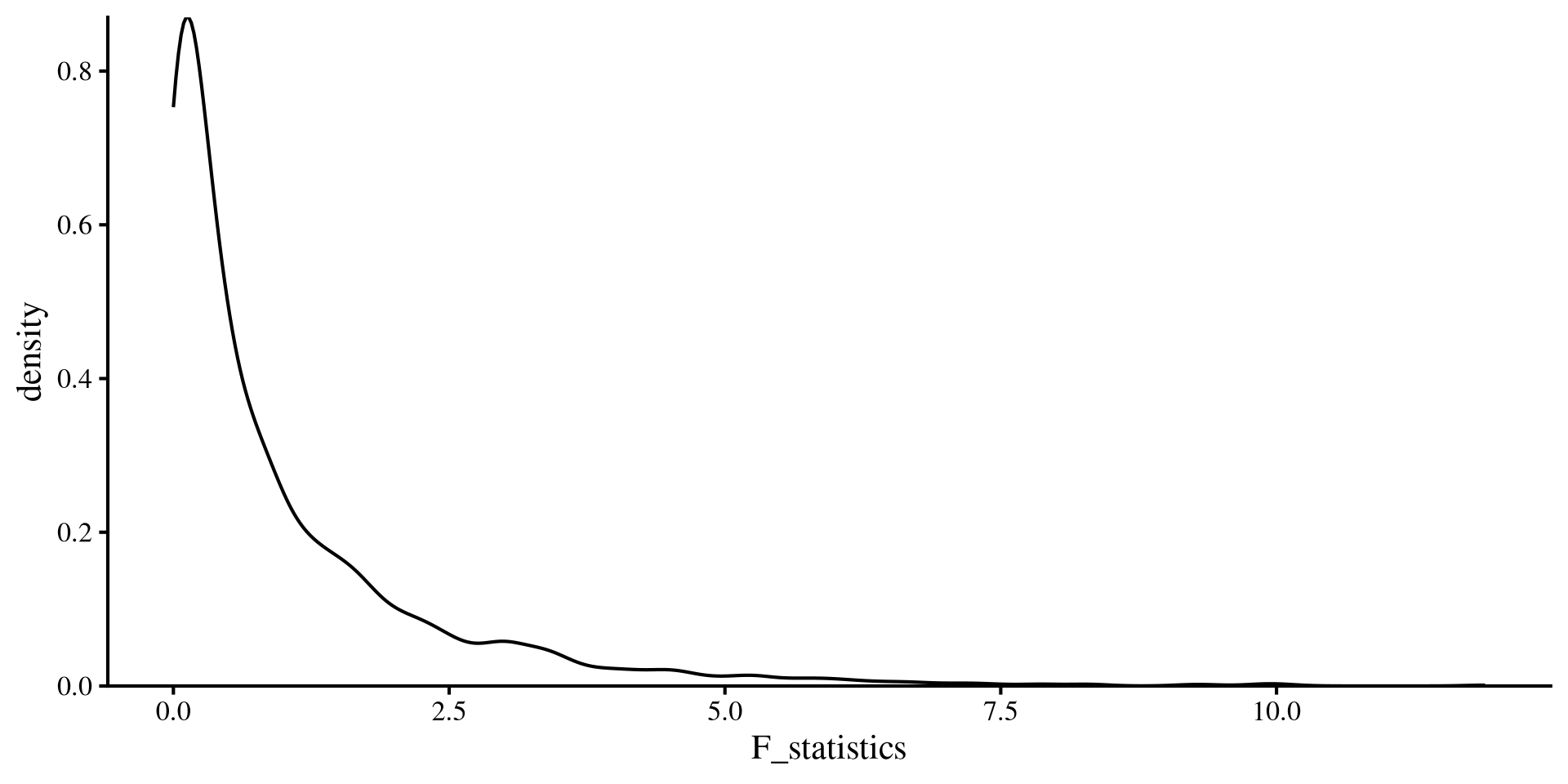
References
Appendix: nice_3D_plot() Showcase
Example with World Happiness Data
You can also add groups to the dots of the 3D plot. Here I am using Region.
There are a bit too many regions, so this is not very practical in this case, but this just an example.
Example with iris Data
A more practical example with the iris dataset, Let’s start with just providing 3 variables.
Name Axes
Let’s name our axes:
Add Groups
Let’s add colors for the 3 different species of flowers:
Add Regression Plane
Let’s add a regression plane:
Pass Arguments to plot_ly()
You can also pass arguments to the plot_ly() function. Let’s make the dots more transparent by passing the opacity = argument to plot_ly():
See here for what “passing arguments” to a function within a function means in R.
Add Interaction Term to Regression Plane
You can also specify an interaction between the two predictors and get an interaction plane. You will see that the reression plane now bends:
See Lab 9 if you are curious to know what is happening.
PSYC 7804 - Lab 4: Introduction To Two-Predictor Regression